
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 15,087,588 – 15,087,689 |
| Length | 101 |
| Max. P | 0.512125 |

| Location | 15,087,588 – 15,087,689 |
|---|---|
| Length | 101 |
| Sequences | 9 |
| Columns | 103 |
| Reading direction | reverse |
| Mean pairwise identity | 64.08 |
| Shannon entropy | 0.73016 |
| G+C content | 0.46949 |
| Mean single sequence MFE | -27.38 |
| Consensus MFE | -9.94 |
| Energy contribution | -9.63 |
| Covariance contribution | -0.31 |
| Combinations/Pair | 1.45 |
| Mean z-score | -1.01 |
| Structure conservation index | 0.36 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.04 |
| SVM RNA-class probability | 0.512125 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 15087588 101 - 24543557 UUGAAAAACAAGCGGUUAAUGUGAAAAACAGGACGAGUUU--AGUUGUUUUCUAUCCGAGGGGACGCGGCGUUGCAACAACAGCGACGCUAUGCGACCGCAUU ...........((((((..(((.....)))..........--....(((..((.....))..)))((((((((((.......))))))))..))))))))... ( -30.40, z-score = -1.58, R) >droSim1.chr3L 14407147 103 - 22553184 UUGAAAAACAAGCGGUUAAUGUGAAAAACAGGACGAGUUUUCGUGUGUUUUCUAUCCGAGAGGACGCGGCGUUGCAACAACAGCGACGCUAUGCGACCGCAUU ...........((((((...(((.((((((.(.((......))).))))))...(((....))))))((((((((.......))))))))....))))))... ( -35.00, z-score = -2.42, R) >droSec1.super_0 7201680 103 - 21120651 UUGAAAAACAAGCGGUUAAUGUGAAAAAGAGGACGAGUUUUCGUGUGUUUUCUAUCCGAGAGGACGCGGAGUUGCAACAACAGCGACGCUAUGCGACCGCAUU ...........((((((.....(((((...(.(((......))).).)))))..(((....))).(((..((((......))))..))).....))))))... ( -30.60, z-score = -1.54, R) >droYak2.chr3L 15180969 101 - 24197627 UUGAAAAACAAGCGGUUAAAGCGCAUAAUAGAACAAGUUU--UGUUGUUUUCUAUUUGAGAGGACGCGGCGUUGCAACAACAGCGACGCUAUGCGACCGCAUU ...........((((((...((((..(((((((....)))--))))(((((((.....)))))))))((((((((.......))))))))..))))))))... ( -34.60, z-score = -3.34, R) >droEre2.scaffold_4784 15044144 101 - 25762168 UUGAAAAACAAGCGCUUAAAGUGGAAAAUAGGACGAGUUU--AGUUGUUUUCUAUCUGAGAGGACGCGGCGUUGCAACAACAGCGACGCUAUGCGACCGCAUU ...........((.((((..(((((((((..(((......--.)))))))))))).)))).((.(((((((((((.......))))))))..))).))))... ( -30.30, z-score = -1.80, R) >droAna3.scaffold_13337 5395254 89 + 23293914 ------------CUAGAAAAGAAGAAAACAAAGCUGGUUUUGUAUCAAUUUCGGU--GUGAGGACGCGGCGCUGCACUGAAAGUGACGCUAUGCGACCGCAUU ------------....................((.((((..(..(((.(((((((--((.(((......).))))))))))).)))..).....))))))... ( -20.30, z-score = 0.52, R) >dp4.chrXR_group6 12663375 84 + 13314419 -------------UAUCAAACGAAGAGAUACAGC-AUAUGUGCAUUAAAGUGAGU--GAGAGGACGCCGCGUUGCAGUGACAGUGACGCUAUG---CUGCAUU -------------.............(((.((((-(((.(((((((...(((.((--(......))))))(((.....))))))).)))))))---))).))) ( -20.50, z-score = -0.11, R) >droPer1.super_12 1081189 84 + 2414086 -------------UAUCAAACGAGGAGAUACAGC-AUAUGUGCAUUAAAGUGAGU--GAGAGGACGCCGCGUUGCAGUGACAGUGACGCUAUG---CUGCAUU -------------.............(((.((((-(((.(((((((...(((.((--(......))))))(((.....))))))).)))))))---))).))) ( -20.50, z-score = 0.10, R) >droWil1.scaffold_181009 654477 101 + 3585778 AACAAAAACAUUGAGUGGAUAAGCGAAACGCAGC-AGUUGU-UCUCGAAUUGGGCACGGAGUACAGUGACGACGCGGCGUCAGCGACGCUAUGCGAUAGCAAU .............((((.....((...((((.((-.(((((-(.....((((..(.....)..)))))))))))).))))..))..)))).(((....))).. ( -24.20, z-score = 1.05, R) >consensus UUGAAAAACAAGCGGUUAAAGUGAAAAACAGAACGAGUUUU_UAUUGUUUUCUAUCCGAGAGGACGCGGCGUUGCAACAACAGCGACGCUAUGCGACCGCAUU .............................................................((.((..(((((((.......)))))))..))...))..... ( -9.94 = -9.63 + -0.31)

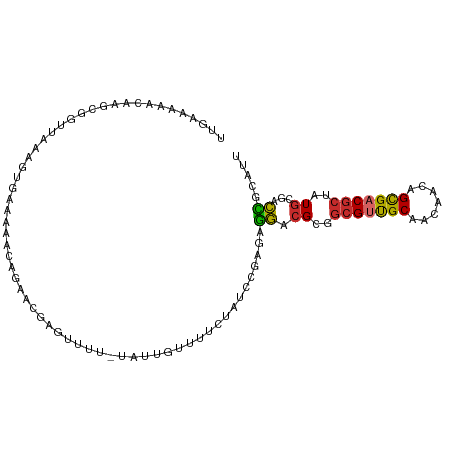
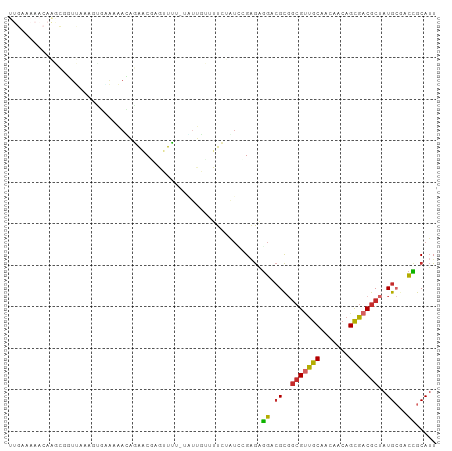
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:27:50 2011