
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 14,953,285 – 14,953,335 |
| Length | 50 |
| Max. P | 0.988289 |

| Location | 14,953,285 – 14,953,335 |
|---|---|
| Length | 50 |
| Sequences | 5 |
| Columns | 52 |
| Reading direction | forward |
| Mean pairwise identity | 87.40 |
| Shannon entropy | 0.22562 |
| G+C content | 0.34523 |
| Mean single sequence MFE | -10.90 |
| Consensus MFE | -8.50 |
| Energy contribution | -9.38 |
| Covariance contribution | 0.88 |
| Combinations/Pair | 1.15 |
| Mean z-score | -2.88 |
| Structure conservation index | 0.78 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.31 |
| SVM RNA-class probability | 0.988289 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
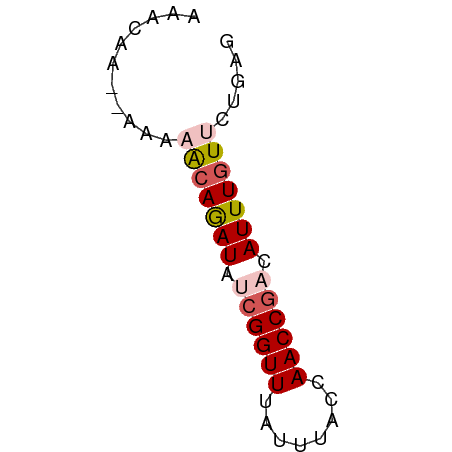
>dm3.chr3L 14953285 50 + 24543557 AAACAA--UAAAGCAGAUAUCGGUUUAUUUACCAACCGACAUUUGUUCUGAG ......--...(((((((.((((((........)))))).)))))))..... ( -12.10, z-score = -3.95, R) >droAna3.scaffold_13337 19128445 52 - 23293914 AAACAAUGGUAAACAAAUCUGGGUUUAUUUACCAACCGUCAUUUGUUGUGAG ......(((((((.((((....)))).)))))))....((((.....)))). ( -10.50, z-score = -0.99, R) >droYak2.chr3L 15040809 50 + 24197627 AAACAA--AAAAAGAGAUAUCGGUUUAUUUACCAACCGCCAUUUGUUUCGAG ......--.....((((((.(((((........))))).....))))))... ( -7.90, z-score = -1.83, R) >droSec1.super_0 7069943 50 + 21120651 AAACAA--AAACACAGAUAUCGGUUUAUUUACCAACCGACAUUUGUUCUGAG ......--....((((((.((((((........)))))).))))))...... ( -11.20, z-score = -3.52, R) >droSim1.chr3L 14270076 50 + 22553184 AAACAA--GAACACAGAUAUCGGUUUAUUUACCAACCGACAUUUGUUCUGAG .....(--(((((..(((.((((((........)))))).)))))))))... ( -12.80, z-score = -4.11, R) >consensus AAACAA__AAAAACAGAUAUCGGUUUAUUUACCAACCGACAUUUGUUCUGAG ...........(((((((.((((((........)))))).)))))))..... ( -8.50 = -9.38 + 0.88)


| Location | 14,953,285 – 14,953,335 |
|---|---|
| Length | 50 |
| Sequences | 5 |
| Columns | 52 |
| Reading direction | reverse |
| Mean pairwise identity | 87.40 |
| Shannon entropy | 0.22562 |
| G+C content | 0.34523 |
| Mean single sequence MFE | -13.10 |
| Consensus MFE | -9.20 |
| Energy contribution | -10.96 |
| Covariance contribution | 1.76 |
| Combinations/Pair | 1.13 |
| Mean z-score | -3.14 |
| Structure conservation index | 0.70 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.28 |
| SVM RNA-class probability | 0.987492 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
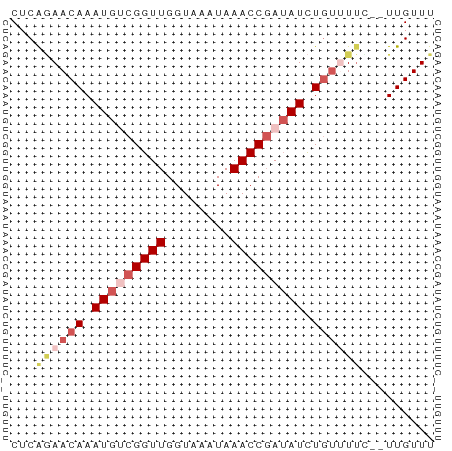
>dm3.chr3L 14953285 50 - 24543557 CUCAGAACAAAUGUCGGUUGGUAAAUAAACCGAUAUCUGCUUUA--UUGUUU ...(((.((.(((((((((........))))))))).)).))).--...... ( -12.20, z-score = -3.02, R) >droAna3.scaffold_13337 19128445 52 + 23293914 CUCACAACAAAUGACGGUUGGUAAAUAAACCCAGAUUUGUUUACCAUUGUUU ............((((((.((((((((((......)))))))))))))))). ( -12.40, z-score = -2.08, R) >droYak2.chr3L 15040809 50 - 24197627 CUCGAAACAAAUGGCGGUUGGUAAAUAAACCGAUAUCUCUUUUU--UUGUUU ....(((((((.((..((((((......))))))....))...)--)))))) ( -9.30, z-score = -1.55, R) >droSec1.super_0 7069943 50 - 21120651 CUCAGAACAAAUGUCGGUUGGUAAAUAAACCGAUAUCUGUGUUU--UUGUUU ...((((((.(((((((((........)))))))))...)))))--)..... ( -14.50, z-score = -3.84, R) >droSim1.chr3L 14270076 50 - 22553184 CUCAGAACAAAUGUCGGUUGGUAAAUAAACCGAUAUCUGUGUUC--UUGUUU ...((((((.(((((((((........)))))))))...)))))--)..... ( -17.10, z-score = -5.19, R) >consensus CUCAGAACAAAUGUCGGUUGGUAAAUAAACCGAUAUCUGUUUUC__UUGUUU ...((((((.(((((((((........))))))))).))))))......... ( -9.20 = -10.96 + 1.76)


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:27:20 2011