| Sequence ID | dm3.chr3L |
|---|---|
| Location | 14,950,171 – 14,950,280 |
| Length | 109 |
| Max. P | 0.997911 |

| Location | 14,950,171 – 14,950,280 |
|---|---|
| Length | 109 |
| Sequences | 9 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 70.69 |
| Shannon entropy | 0.59597 |
| G+C content | 0.44452 |
| Mean single sequence MFE | -33.42 |
| Consensus MFE | -12.06 |
| Energy contribution | -12.47 |
| Covariance contribution | 0.41 |
| Combinations/Pair | 1.50 |
| Mean z-score | -3.27 |
| Structure conservation index | 0.36 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 3.21 |
| SVM RNA-class probability | 0.997911 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
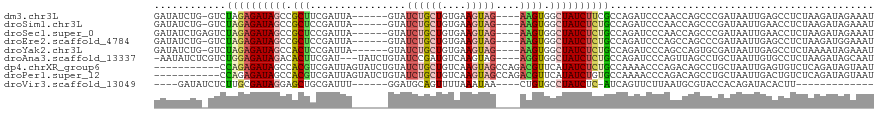
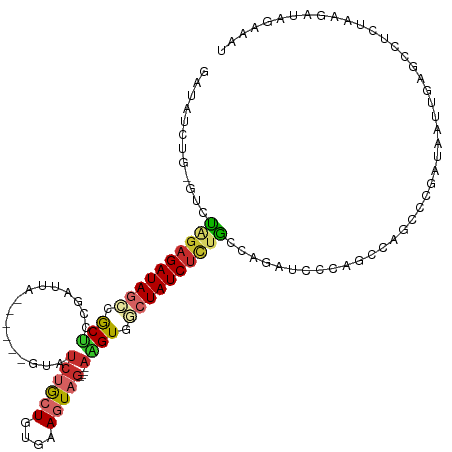
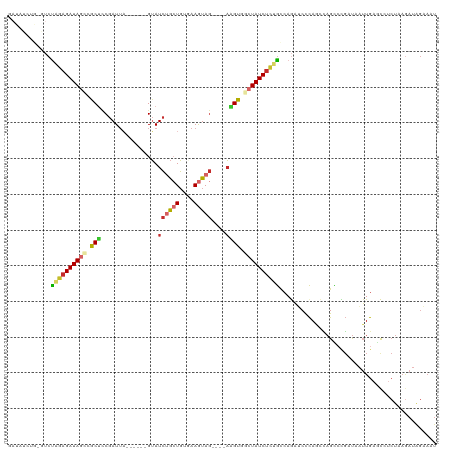
>dm3.chr3L 14950171 109 + 24543557 GAUAUCUG-GUCUAGAGAUAGCCGCUUCGAUUA------GUAUCUGCUGUGAAGUAG----AAGUGGCUAUCUUCGCCAGAUCCCAACCAGCCCGAUAAUUGAGCCUCUAAGAUAGAAAU (..(((((-((...(((((((((((((((((..------..)))((((....)))))----))))))))))))).)))))))..).....((.(((...))).))............... ( -35.80, z-score = -3.61, R) >droSim1.chr3L 14266930 109 + 22553184 GAUAUCUG-GUCUAGAGAUAGCCGCUCCGAUUA------GUAUCUGCUGUGAAGUAG----AAGUGGCUAUCUCUGCCAGAUCCCAACCAGCCCGAUAAUUGAACCUCUAAGAUAGAAAU (..(((((-((..(((((((((((((.......------...((((((....)))))----)))))))))))))))))))))..)................................... ( -36.10, z-score = -4.22, R) >droSec1.super_0 7066920 110 + 21120651 GAUAUCUGAGUCUAGAGAUAGCCGCUCCGAUUA------GUAUCUGCUGUGAAGUAG----AAGUGGCUAUCUCUGCCAGAUCCCAACCAGCCCGAUAAUUGAACCUCUAAGAUAGAAAU (..(((((.((..(((((((((((((.......------...((((((....)))))----)))))))))))))))))))))..)................................... ( -32.30, z-score = -3.41, R) >droEre2.scaffold_4784 14907792 109 + 25762168 GAUAUCUG-GUCUAGAGAUAGCCGCUCCGAUUA------GUAUCUGCUGUGAAGUAG----AAGUGGCUAUCUCUGCCAGAUCCCAGCCAGCCCGAUAAUUGAGCCUCUAAGAUGGAAAU ...(((((-((..(((((((((((((.......------...((((((....)))))----))))))))))))))))))))).(((....((.(((...))).))........))).... ( -37.80, z-score = -3.26, R) >droYak2.chr3L 15037843 109 + 24197627 GAUAUCUG-GUCUAGAGAUAGCCACUCCGAUUA------GUAUCUGCUGUGAAGUAG----AAGUGGCUAUCUCUGCCAGAUCCCAGCCAGUGCGAUAAUUGAGCCUCUAAAAUAGAAAU (..(((((-((..(((((((((((((.......------...((((((....)))))----)))))))))))))))))))))..).((((((......)))).))............... ( -40.50, z-score = -5.01, R) >droAna3.scaffold_13337 19125451 112 - 23293914 -AAUAUCUCGUCUGGAGAUAGACACUUCGAU---UAUCUGUAUCCGAUGUCAAGUAG----AGGUGGCUAUCUCUGCCAGAUCCCAGUUAGCCUGCUAAUUGUGCCUCUAAGAUAGCAAU -..(((((((((.(((.(((((.........---..))))).))))))).....(((----(((((((.......)))......(((((((....))))))).))))))))))))..... ( -32.30, z-score = -1.72, R) >dp4.chrXR_group6 12528927 109 - 13314419 -----------CCAGAGAUAGCCACGUCGAUUAGUAUCUGUAUCUGCUGUCAAGUAGCCAGACGUUCAUAUCUCUGCCAAAACCCAGACAGCCUGCUAAUUGAGUGUCUCAGAUAGUAAU -----------.((((((((...((((((((....))).....(((((....)))))...)))))...)))))))).................(((((.(((((...))))).))))).. ( -29.70, z-score = -2.72, R) >droPer1.super_12 946055 109 - 2414086 -----------CCAGAGAUAGCCACGUCGAUUAGUAUCUGUAUCUGCUGUCAAGUAGCCAGACGUUCAUAUCUGUGCCAAAACCCAGACAGCCUGCUAAUUGACUGUCUCAGAUAGUAAU -----------...((((((.....(((((((((((.((((...((((....))))((((((........)))).))..........))))..))))))))))))))))).......... ( -29.10, z-score = -2.38, R) >droVir3.scaffold_13049 7453007 92 - 25233164 ----GAUAUCUCUUGCGAUAGGAGCUGCGAUUU------GGAUGCAGUUUUAAAUAA----CUGUGCCUAUCUC-AUCAGUUCUUAAUGCGUACCACAGAUACACUU------------- ----..(((((..((((.(((((((((.((..(------((..((((((......))----))))..)))..))-..))))))))).))))......))))).....------------- ( -27.20, z-score = -3.12, R) >consensus GAUAUCUG_GUCUAGAGAUAGCCGCUCCGAUUA______GUAUCUGCUGUGAAGUAG____AAGUGGCUAUCUCUGCCAGAUCCCAGCCAGCCCGAUAAUUGAGCCUCUAAGAUAGAAAU ............((((((((((.(((.................(((((....))))).....))).))))))))))............................................ (-12.06 = -12.47 + 0.41)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:27:19 2011