| Sequence ID | dm3.chr3L |
|---|---|
| Location | 14,654,151 – 14,654,241 |
| Length | 90 |
| Max. P | 0.950165 |

| Location | 14,654,151 – 14,654,241 |
|---|---|
| Length | 90 |
| Sequences | 9 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 64.83 |
| Shannon entropy | 0.62890 |
| G+C content | 0.46469 |
| Mean single sequence MFE | -18.49 |
| Consensus MFE | -6.93 |
| Energy contribution | -7.01 |
| Covariance contribution | 0.08 |
| Combinations/Pair | 1.30 |
| Mean z-score | -2.03 |
| Structure conservation index | 0.38 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.56 |
| SVM RNA-class probability | 0.950165 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
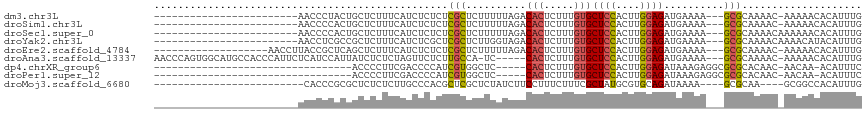
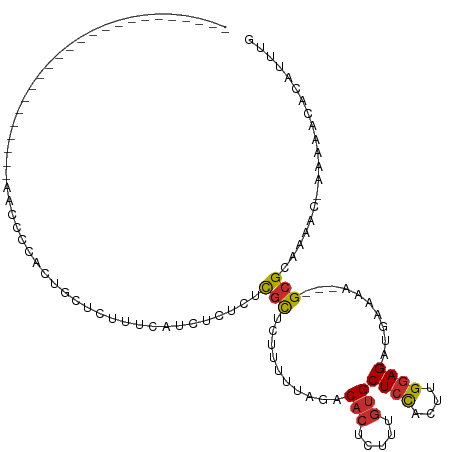
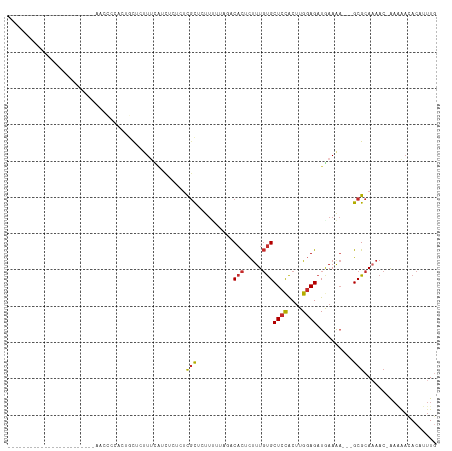
>dm3.chr3L 14654151 90 - 24543557 ------------------------AACCCUACUGCUCUUUCAUCUCUCUCGCUCUUUUUAGACACUCUUUGUGCUCCACUUGGAGAUGAAAA---GCGCAAAAC-AAAAACACAUUUG ------------------------.........(((.((((((((((.....(((....)))(((.....)))........)))))))))))---)).......-............. ( -17.30, z-score = -2.82, R) >droSim1.chr3L 13984910 90 - 22553184 ------------------------AACCCCACUGCUCUUUCAUCUCUCUCGCUCUUUUUAGACACUCUUUGUGCUCCACUUGGAGAUGAAAA---GCGCAAAAC-AAAAACACAUUUG ------------------------.........(((.((((((((((.....(((....)))(((.....)))........)))))))))))---)).......-............. ( -17.30, z-score = -2.71, R) >droSec1.super_0 6779413 91 - 21120651 ------------------------AACCCCACUGCUCUUUCAUCUCUCUCGCUCUUUUUAGACACUCUUUGUGCUCCACUUGGAGAUGAAAA---GCGCAAAACAAAAAACACAUUUG ------------------------.........(((.((((((((((.....(((....)))(((.....)))........)))))))))))---))..................... ( -17.30, z-score = -2.66, R) >droYak2.chr3L 14743351 91 - 24197627 ------------------------AACCUCGCCGCUCUUUCAUCUCGCUCGCUCUUGGUAGACACUCUUUGUGCUCCACUUGGAGAUGAAAA---GCGCAAAACAAAACAUACAUUUG ------------------------......((.(((.((((((((((....)...(((.((.(((.....))))))))....))))))))))---))))................... ( -22.90, z-score = -2.90, R) >droEre2.scaffold_4784 14624532 95 - 25762168 -------------------AACCUUACCGCUCAGCUCUUUCAUCUCUCUCGCUCUUUUUAGACACUCUUUGUGCUCCACUUGGAGAUGAAAA---GCGCAAAAC-AAAAACACAUUUG -------------------.........((...(((.((((((((((.....(((....)))(((.....)))........)))))))))))---)))).....-............. ( -17.80, z-score = -2.05, R) >droAna3.scaffold_13337 18847693 108 + 23293914 AACCCAGUGGCAUGCCACCCAUUCUCAUCCAUUAUCUCUCUAGUUCUCUUGCCA-UC-----CACUCUUUGUGCUCCACUUGGAGAUGAAAA---GCGCAAAAC-AAAAACACAUUUG ......((((....))))........................(((...(((((.-..-----(((.....)))((((....)))).......---).)))))))-............. ( -16.10, z-score = -0.18, R) >dp4.chrXR_group6 12223609 78 + 13314419 ---------------------------------ACCCCUUCGACCCCAUCGUGGCUC-----CACUCUUUGUGCUCCACUUGGAGAUAAAGAGGCGCGCACAAC-AACAA-ACAUUUC ---------------------------------.................(((((..-----..((((((((.((((....))))))))))))..)).)))...-.....-....... ( -20.70, z-score = -2.56, R) >droPer1.super_12 635357 78 + 2414086 ---------------------------------ACCCCUUCGACCCCAUCGUGGCUC-----CACUCUUUGUGCUCCACUUGGAGAUAAAGAGGCGCGCACAAC-AACAA-ACAUUUC ---------------------------------.................(((((..-----..((((((((.((((....))))))))))))..)).)))...-.....-....... ( -20.70, z-score = -2.56, R) >droMoj3.scaffold_6680 4812493 85 + 24764193 -------------------------CACCCGCGCUCUCUCUUGCCCACGCUCGCUCUAUCUUCCUUUCUUUCGCUAUGCGUGCAGAUAAAA----GCGCAA----GCGGCCACAUUUG -------------------------.....(((((.....((((..((((..((..................))...)))))))).....)----))))..----............. ( -16.27, z-score = 0.16, R) >consensus ________________________AACCCCACUGCUCUUUCAUCUCUCUCGCUCUUUUUAGACACUCUUUGUGCUCCACUUGGAGAUGAAAA___GCGCAAAAC_AAAAACACAUUUG .................................................(((..........(((.....)))((((....))))..........))).................... ( -6.93 = -7.01 + 0.08)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:26:24 2011