
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 14,636,744 – 14,636,825 |
| Length | 81 |
| Max. P | 0.607030 |

| Location | 14,636,744 – 14,636,825 |
|---|---|
| Length | 81 |
| Sequences | 12 |
| Columns | 90 |
| Reading direction | forward |
| Mean pairwise identity | 74.62 |
| Shannon entropy | 0.52282 |
| G+C content | 0.41723 |
| Mean single sequence MFE | -11.30 |
| Consensus MFE | -6.26 |
| Energy contribution | -5.91 |
| Covariance contribution | -0.35 |
| Combinations/Pair | 1.25 |
| Mean z-score | -1.05 |
| Structure conservation index | 0.55 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.24 |
| SVM RNA-class probability | 0.607030 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 14636744 81 + 24543557 ---AGUCAAUUGACGCUUUGUUUUAUAACUUCCA-AGACGUAAGACAAUAAACAACGA---GUCUUUCGCUUACCACA--GCCACUCCCC ---.(((....)))...(((((((((..((....-))..)))))))))........((---((.....(((......)--)).))))... ( -11.70, z-score = -1.30, R) >droSim1.chr3L 13967405 81 + 22553184 ---AGUCAAUUGACGCUUUGUUUUAUAACUUCCA-AGACGUAAGACAAUAAACAACGA---GUCUUUCGCUUACCGCA--GCCACUCCCC ---.(((....)))...(((((((((..((....-))..)))))))))........((---((..((.((.....)))--)..))))... ( -11.80, z-score = -0.88, R) >droSec1.super_0 6762168 81 + 21120651 ---AGUCAAUUGACGCUUUGUUUUAUAACUUCCA-AGACGUAAGACAAUAAACAACGA---GUCUUUCGCUUACCGCA--GCCACUCCCC ---.(((....)))...(((((((((..((....-))..)))))))))........((---((..((.((.....)))--)..))))... ( -11.80, z-score = -0.88, R) >droYak2.chr3L 14725347 81 + 24197627 ---AGUCAAUUGACGCUUUGUUUUAUAACUUCCA-AGACGUAAGACAAUAAACAACGA---GUCUUUCGCUUACCGCA--GCCACUCCCC ---.(((....)))...(((((((((..((....-))..)))))))))........((---((..((.((.....)))--)..))))... ( -11.80, z-score = -0.88, R) >droEre2.scaffold_4784 14606998 81 + 25762168 ---AGUCAAUUGACGCUUUGUUUUAUAACUUCCA-AGACGUAAGACAAUAAACAACGA---GUCUUUCGCUUGCCGCA--GCCACUCCCC ---.(((....)))((...(((....)))...((-((.((.(((((............---))))).))))))..)).--.......... ( -13.60, z-score = -1.03, R) >droAna3.scaffold_13337 18832652 86 - 23293914 ---AGUCAAUUGACGCUUUGUUUUAUAACUUCCA-AGACGUAAGACAAUAAACAACAACUCGUCUCUCGCUUACCACACAGCCACACCCC ---.(((....)))(((..(((....)))....(-((.((..((((...............))))..))))).......)))........ ( -10.16, z-score = -1.07, R) >dp4.chrXR_group6 1106710 77 + 13314419 AGUCGUCAAUUGACGCUUUGUUUUAUAACUUCCC-AGUCAUAAGACAAUAAACAACAG-----GUCUCCAUGGCCGCC--GCCCC----- .((((.....))))((.(((((((((.(((....-))).))))))))).........(-----(((.....)))))).--.....----- ( -18.20, z-score = -2.98, R) >droPer1.super_12 1795014 77 + 2414086 AGUCGUCAAUUGACGCUUUGUUUUAUAACUUCCC-AGUCAUAAGACAAUAAACAACAG-----GUCUCCAUGGCCGCC--GCCCC----- .((((.....))))((.(((((((((.(((....-))).))))))))).........(-----(((.....)))))).--.....----- ( -18.20, z-score = -2.98, R) >droWil1.scaffold_181136 937058 79 - 2313701 ---AAUCAAUUGACGCUUUGUUUUAUAGCUUCCA-ACUCGUAAAAUAAUGAACAACAA-----GUCUCGCUUAUUCUU--UCCAUUUCCC ---........((((((.........))).....-..((((......)))).......-----)))............--.......... ( -5.20, z-score = 0.11, R) >droVir3.scaffold_13049 8504315 79 - 25233164 ---AGUCAAUUGACGCUUUGUUUUAUAACUUGCA-AGCCAUAAGAUAAUAAACAACAUAUUUCAUCUCCCUUGCUCCCUCACU------- ---.(((....))).................(((-((.....((((((((.......))))..))))..))))).........------- ( -7.50, z-score = -0.66, R) >droMoj3.scaffold_6680 19971794 79 + 24764193 ---AGUCAAUUGACGCUUUGUUUUAUAACCUGCAUAGCCGUAAGAUAAUAAACAAC-UCCUCCCUCUCACUCGCUCUCUCCCU------- ---.(((....)))(((.(((..........))).)))..................-..........................------- ( -6.70, z-score = -0.46, R) >droGri2.scaffold_15110 16375326 79 + 24565398 ---AGUCAAUUGACGCUUUGUUUUAUAAAUUUCA-AGCCAGAAGAUAAUAAACAACAGUUUCGAACAUGCUCUCUCACUCGCU------- ---......((((.(((((((((......((((.-.....)))).....)))))).))).))))...................------- ( -8.90, z-score = 0.49, R) >consensus ___AGUCAAUUGACGCUUUGUUUUAUAACUUCCA_AGACGUAAGACAAUAAACAACAA___GUCUCUCGCUUACCGCA__GCCACUCCCC ....(((....)))...(((((((((.............))))))))).......................................... ( -6.26 = -5.91 + -0.35)

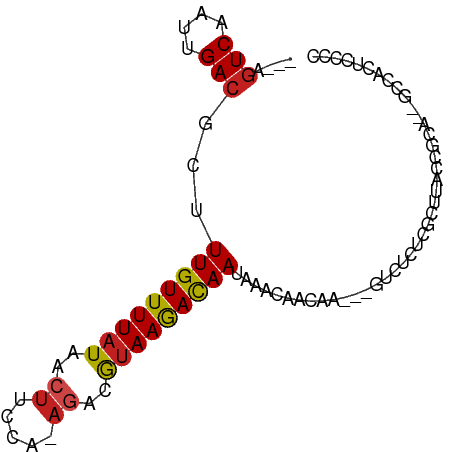
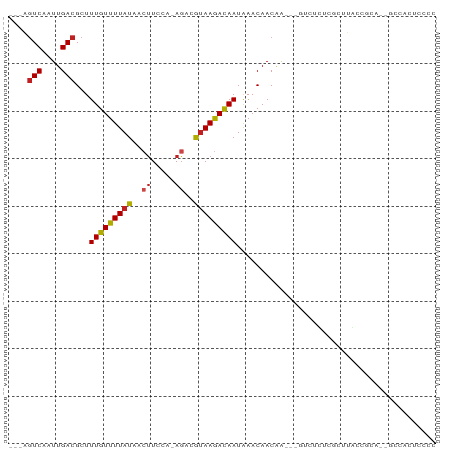
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:26:21 2011