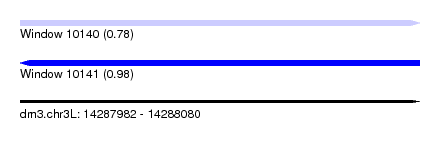

| Sequence ID | dm3.chr3L |
|---|---|
| Location | 14,287,982 – 14,288,080 |
| Length | 98 |
| Max. P | 0.981762 |

| Location | 14,287,982 – 14,288,080 |
|---|---|
| Length | 98 |
| Sequences | 8 |
| Columns | 113 |
| Reading direction | forward |
| Mean pairwise identity | 68.48 |
| Shannon entropy | 0.62256 |
| G+C content | 0.40474 |
| Mean single sequence MFE | -23.53 |
| Consensus MFE | -10.77 |
| Energy contribution | -11.78 |
| Covariance contribution | 1.00 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.19 |
| Structure conservation index | 0.46 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.68 |
| SVM RNA-class probability | 0.784667 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
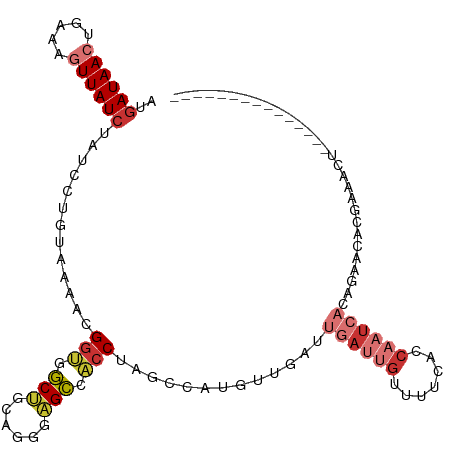
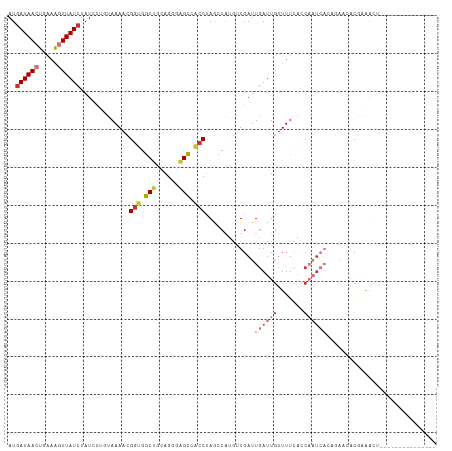
>dm3.chr3L 14287982 98 + 24543557 AUGAUAACUGAAAGUUAUCUAUAUUGUAAAACGGUGGCUGCAGGGAGCCACCUAGCCAUGUUGAUUGAUUGUUUUAACCAAUCACAGAACACGAAGUU--------------- ..((((((.....))))))....((((.....(((((((......))))))).......(((...((((((.......))))))...)))))))....--------------- ( -24.90, z-score = -1.93, R) >droSim1.chr3L 13639603 98 + 22553184 AUGAUAACUGAAAGUUAUCUAUUCUGUAAAACGGUGGCUGCAGGGAGCCACCUAGCCAUGUUGAUUGAUUGUUUUCACCAAUCACAGAACACGAAGCU--------------- ..((((((.....))))))..((((((.....(((((((......)))))))..............(((((.......))))))))))).........--------------- ( -27.60, z-score = -2.21, R) >droSec1.super_0 6430548 98 + 21120651 AUUAUAACUGAAAGUUAUCUAUUCUGUAAAACGGUGGCUGCAGGGAGCCACCUAGCCAUGUUGAUUGAUUGUUUUCACCAAUCACAGAACACGAAGCU--------------- ...(((((.....)))))...((((((.....(((((((......)))))))..............(((((.......))))))))))).........--------------- ( -23.80, z-score = -1.21, R) >droYak2.chr3L 14382278 98 + 24197627 AUGAUAACUGAAAGUUAUCUAUUCUGUAAAACGGUGGCUGCAGGGAGCUACCUAGCCAUGUUGAUUAAUUGUUUUC---------------CCCAAUCACAGAACACGAAGCU ..((((((.....))))))..((((((..((((.(((((..(((......))))))))))))((((..........---------------...))))))))))......... ( -22.82, z-score = -1.00, R) >droEre2.scaffold_4784 14274982 98 + 25762168 AUGAUAACUGCAAGUUAUCUAUACUGUAAAAUGUUGGCUGCAGGGAGCCACCUAGCCAUGUUGAUUGAUUGUUUUCACCAAUCACAGAACACGAAGCU--------------- ..((((((.....))))))....((((.......(((((..(((......))))))))........(((((.......)))))))))...........--------------- ( -22.20, z-score = -0.43, R) >droAna3.scaffold_13337 18521148 111 - 23293914 AUGAUAACCGAGGGUUAUCUAUCCUUGAAAACGGUGGCAGCAGGGAGC--CCUAAUCGAAUUGAUUGAUUGCUUUUUUCCUUGUCUUUUAUCCCAAUCACAGAAAACAGAGUU .((((...(((((((.....))))))).....(((((.(((((((((.--..(((((((.....))))))).....))))))).)).)))))...)))).............. ( -24.10, z-score = 0.20, R) >dp4.chrXR_group6 6971125 106 + 13314419 AUGAUAACUGAAACUUAUCUAUCCC-UAAAAUGGCGCCCA-AGAGGGGAGCCAAAGCACACUGAUUGAUUGUUUUUGCCAAUCAACAAUCACAGAACAACACCAAAUA----- ..(((((.......)))))......-.....((((.(((.-....))).)))).......(((..((((((((..(....)..)))))))))))..............----- ( -21.40, z-score = -1.53, R) >droPer1.super_41 332382 106 + 728894 AUGAUAACUGAAACUUAUCUAUCCC-CAAAAUGGCGCCCA-AGAGGGGAGCCAAAGCACACUGAUUGAUUGUUUUUGCCAAUCAACAAUCACAGAACAACACCAAAUA----- ..(((((.......)))))......-.....((((.(((.-....))).)))).......(((..((((((((..(....)..)))))))))))..............----- ( -21.40, z-score = -1.42, R) >consensus AUGAUAACUGAAAGUUAUCUAUCCUGUAAAACGGUGGCUGCAGGGAGCCACCUAGCCAUGUUGAUUGAUUGUUUUCACCAAUCACAGAACACGAAACU_______________ ..((((((.....)))))).............(((.(((......))).))).............((((((.......))))))............................. (-10.77 = -11.78 + 1.00)
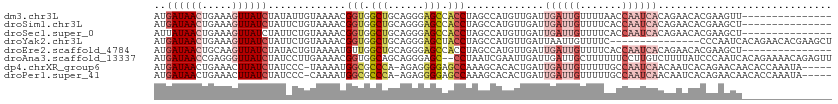

| Location | 14,287,982 – 14,288,080 |
|---|---|
| Length | 98 |
| Sequences | 8 |
| Columns | 113 |
| Reading direction | reverse |
| Mean pairwise identity | 68.48 |
| Shannon entropy | 0.62256 |
| G+C content | 0.40474 |
| Mean single sequence MFE | -27.72 |
| Consensus MFE | -12.48 |
| Energy contribution | -13.47 |
| Covariance contribution | 1.00 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.96 |
| Structure conservation index | 0.45 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.08 |
| SVM RNA-class probability | 0.981762 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
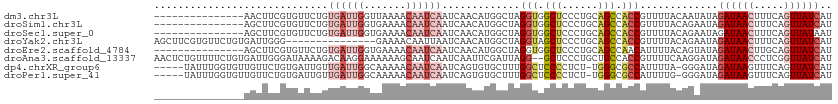
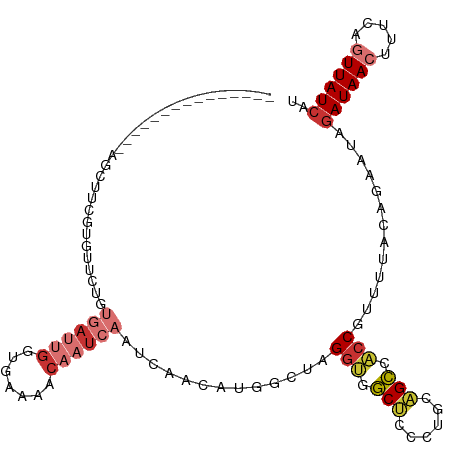
>dm3.chr3L 14287982 98 - 24543557 ---------------AACUUCGUGUUCUGUGAUUGGUUAAAACAAUCAAUCAACAUGGCUAGGUGGCUCCCUGCAGCCACCGUUUUACAAUAUAGAUAACUUUCAGUUAUCAU ---------------....(((((((...((((((.......))))))...)))))))...(((((((......))))))).............((((((.....)))))).. ( -31.40, z-score = -4.82, R) >droSim1.chr3L 13639603 98 - 22553184 ---------------AGCUUCGUGUUCUGUGAUUGGUGAAAACAAUCAAUCAACAUGGCUAGGUGGCUCCCUGCAGCCACCGUUUUACAGAAUAGAUAACUUUCAGUUAUCAU ---------------.......(((((((((((((((.......)))))))..........(((((((......))))))).....))))))))((((((.....)))))).. ( -33.90, z-score = -4.26, R) >droSec1.super_0 6430548 98 - 21120651 ---------------AGCUUCGUGUUCUGUGAUUGGUGAAAACAAUCAAUCAACAUGGCUAGGUGGCUCCCUGCAGCCACCGUUUUACAGAAUAGAUAACUUUCAGUUAUAAU ---------------.......(((((((((((((((.......)))))))..........(((((((......))))))).....)))))))).(((((.....)))))... ( -30.20, z-score = -3.20, R) >droYak2.chr3L 14382278 98 - 24197627 AGCUUCGUGUUCUGUGAUUGGG---------------GAAAACAAUUAAUCAACAUGGCUAGGUAGCUCCCUGCAGCCACCGUUUUACAGAAUAGAUAACUUUCAGUUAUCAU .......((((((((((.((((---------------.....)............(((((..((((....))))))))))))..))))))))))((((((.....)))))).. ( -27.10, z-score = -2.17, R) >droEre2.scaffold_4784 14274982 98 - 25762168 ---------------AGCUUCGUGUUCUGUGAUUGGUGAAAACAAUCAAUCAACAUGGCUAGGUGGCUCCCUGCAGCCAACAUUUUACAGUAUAGAUAACUUGCAGUUAUCAU ---------------(((..((((((...((((((.......))))))...))))))))).(((.((.....)).)))................((((((.....)))))).. ( -26.00, z-score = -1.39, R) >droAna3.scaffold_13337 18521148 111 + 23293914 AACUCUGUUUUCUGUGAUUGGGAUAAAAGACAAGGAAAAAAGCAAUCAAUCAAUUCGAUUAGG--GCUCCCUGCUGCCACCGUUUUCAAGGAUAGAUAACCCUCGGUUAUCAU .....((((((.(((.......))).)))))).((.....(((....((((.....))))(((--....)))))).)).((........))...(((((((...))))))).. ( -21.10, z-score = 0.90, R) >dp4.chrXR_group6 6971125 106 - 13314419 -----UAUUUGGUGUUGUUCUGUGAUUGUUGAUUGGCAAAAACAAUCAAUCAGUGUGCUUUGGCUCCCCUCU-UGGGCGCCAUUUUA-GGGAUAGAUAAGUUUCAGUUAUCAU -----....(((((.((((((((((((((((((((.......))))))).)))).)))..((((.(((....-.))).)))).....-))))))((......))...))))). ( -26.00, z-score = -0.50, R) >droPer1.super_41 332382 106 - 728894 -----UAUUUGGUGUUGUUCUGUGAUUGUUGAUUGGCAAAAACAAUCAAUCAGUGUGCUUUGGCUCCCCUCU-UGGGCGCCAUUUUG-GGGAUAGAUAAGUUUCAGUUAUCAU -----....(((((.((((((((((((((((((((.......))))))).)))).)))..((((.(((....-.))).)))).....-))))))((......))...))))). ( -26.10, z-score = -0.27, R) >consensus _______________AGCUUCGUGUUCUGUGAUUGGUGAAAACAAUCAAUCAACAUGGCUAGGUGGCUCCCUGCAGCCACCGUUUUACAGAAUAGAUAACUUUCAGUUAUCAU .............................((((((.......)))))).............(((.(((......))).))).............((((((.....)))))).. (-12.48 = -13.47 + 1.00)
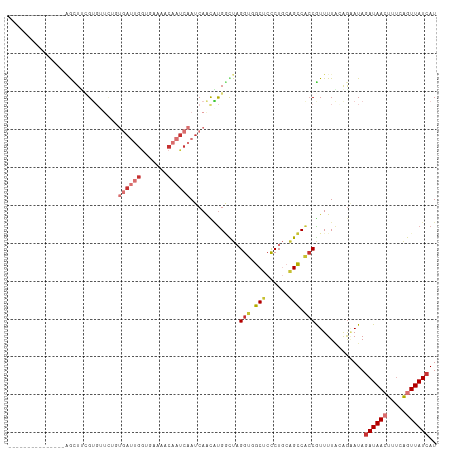
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:25:32 2011