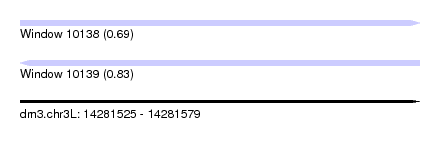

| Sequence ID | dm3.chr3L |
|---|---|
| Location | 14,281,525 – 14,281,579 |
| Length | 54 |
| Max. P | 0.830427 |

| Location | 14,281,525 – 14,281,579 |
|---|---|
| Length | 54 |
| Sequences | 6 |
| Columns | 61 |
| Reading direction | forward |
| Mean pairwise identity | 83.55 |
| Shannon entropy | 0.29758 |
| G+C content | 0.43691 |
| Mean single sequence MFE | -10.28 |
| Consensus MFE | -7.72 |
| Energy contribution | -7.83 |
| Covariance contribution | 0.11 |
| Combinations/Pair | 1.15 |
| Mean z-score | -1.53 |
| Structure conservation index | 0.75 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.41 |
| SVM RNA-class probability | 0.685938 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
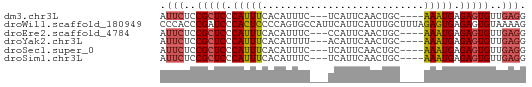
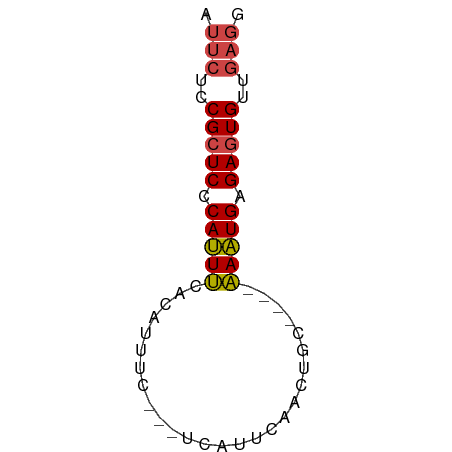
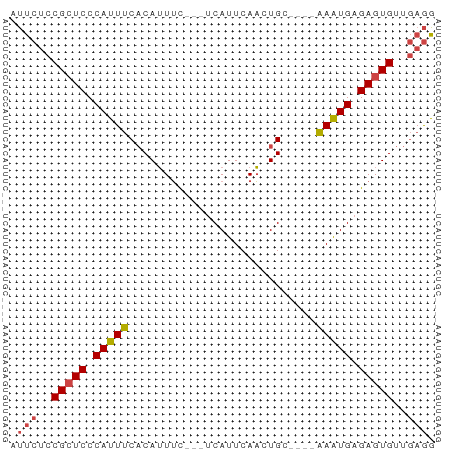
>dm3.chr3L 14281525 54 + 24543557 AUUCUCCGCUCCCAUUUCACAUUUC---UCAUUCAACUGC----AAAUGAGAGUGUUGAGG .(((..(((((.(((((((......---.........)).----))))).)))))..))). ( -10.56, z-score = -1.41, R) >droWil1.scaffold_180949 3577008 61 + 6375548 CCCACCCGAUCCCACUCCCCAGUGCCAUUCAUUCAUUUGCUUUAGAGUGAGAGUGUAAAAG ............((((....)))).(((((..((((((......)))))))))))...... ( -8.90, z-score = -1.66, R) >droEre2.scaffold_4784 14268533 54 + 25762168 AUUCUCCGCUCCCAUUUCACAUUUC---CCAUUCAACUGC----AAAUGAGAGUGUUGAGG .(((..(((((.(((((((......---.........)).----))))).)))))..))). ( -10.56, z-score = -1.62, R) >droYak2.chr3L 14375737 54 + 24197627 AUUCUCCGCUCCCAUUUCACAUUUU---ACAUUCAACUGC----AAAUGAGAGUGUUGAGG .(((..(((((.(((((((......---.........)).----))))).)))))..))). ( -10.56, z-score = -1.67, R) >droSec1.super_0 6424230 54 + 21120651 AUUCUCCGCUCCCAUUUCACAUUUC---UCAUUCAACUGC----AAAUGAGAGUGUUGAGG .(((..(((((.(((((((......---.........)).----))))).)))))..))). ( -10.56, z-score = -1.41, R) >droSim1.chr3L 13633316 54 + 22553184 AUUCUCCGCUCCCAUUUCACAUUUC---UCAUUCAACUGC----AAAUGAGAGUGUUGAGG .(((..(((((.(((((((......---.........)).----))))).)))))..))). ( -10.56, z-score = -1.41, R) >consensus AUUCUCCGCUCCCAUUUCACAUUUC___UCAUUCAACUGC____AAAUGAGAGUGUUGAGG .(((..(((((.(((((...........................))))).)))))..))). ( -7.72 = -7.83 + 0.11)
| Location | 14,281,525 – 14,281,579 |
|---|---|
| Length | 54 |
| Sequences | 6 |
| Columns | 61 |
| Reading direction | reverse |
| Mean pairwise identity | 83.55 |
| Shannon entropy | 0.29758 |
| G+C content | 0.43691 |
| Mean single sequence MFE | -13.60 |
| Consensus MFE | -11.19 |
| Energy contribution | -10.86 |
| Covariance contribution | -0.33 |
| Combinations/Pair | 1.27 |
| Mean z-score | -1.50 |
| Structure conservation index | 0.82 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.83 |
| SVM RNA-class probability | 0.830427 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
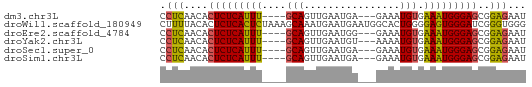
>dm3.chr3L 14281525 54 - 24543557 CCUCAACACUCUCAUUU----GCAGUUGAAUGA---GAAAUGUGAAAUGGGAGCGGAGAAU .(((....(((((((((----(((.........---....))).)))))))))..)))... ( -13.22, z-score = -1.48, R) >droWil1.scaffold_180949 3577008 61 - 6375548 CUUUUACACUCUCACUCUAAAGCAAAUGAAUGAAUGGCACUGGGGAGUGGGAUCGGGUGGG ......((((((((((((..((...((......))....))..)))))))))....))).. ( -15.10, z-score = -1.33, R) >droEre2.scaffold_4784 14268533 54 - 25762168 CCUCAACACUCUCAUUU----GCAGUUGAAUGG---GAAAUGUGAAAUGGGAGCGGAGAAU .(((....(((((((((----(((.........---....))).)))))))))..)))... ( -13.22, z-score = -1.28, R) >droYak2.chr3L 14375737 54 - 24197627 CCUCAACACUCUCAUUU----GCAGUUGAAUGU---AAAAUGUGAAAUGGGAGCGGAGAAU .(((....(((((((((----(((.((......---.)).))).)))))))))..)))... ( -13.60, z-score = -1.93, R) >droSec1.super_0 6424230 54 - 21120651 CCUCAACACUCUCAUUU----GCAGUUGAAUGA---GAAAUGUGAAAUGGGAGCGGAGAAU .(((....(((((((((----(((.........---....))).)))))))))..)))... ( -13.22, z-score = -1.48, R) >droSim1.chr3L 13633316 54 - 22553184 CCUCAACACUCUCAUUU----GCAGUUGAAUGA---GAAAUGUGAAAUGGGAGCGGAGAAU .(((....(((((((((----(((.........---....))).)))))))))..)))... ( -13.22, z-score = -1.48, R) >consensus CCUCAACACUCUCAUUU____GCAGUUGAAUGA___GAAAUGUGAAAUGGGAGCGGAGAAU .(((....(((((((((....(((................))).)))))))))..)))... (-11.19 = -10.86 + -0.33)
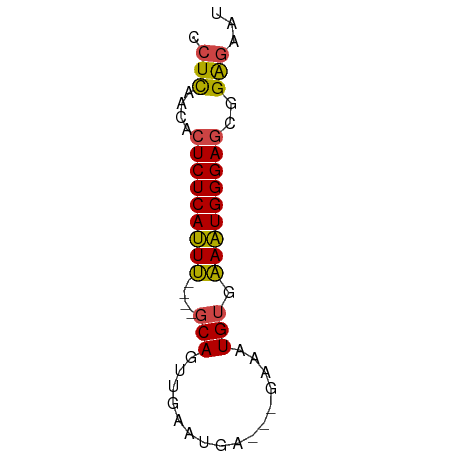
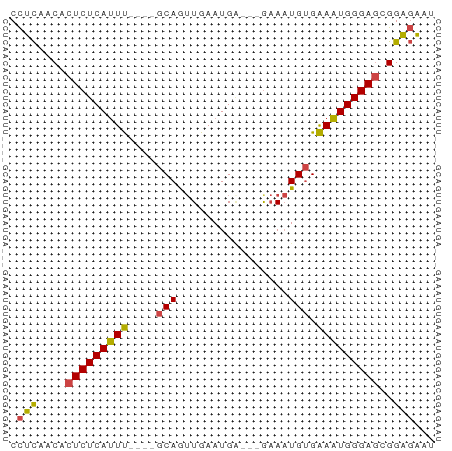
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:25:30 2011