
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 14,267,909 – 14,268,037 |
| Length | 128 |
| Max. P | 0.529558 |

| Location | 14,267,909 – 14,268,037 |
|---|---|
| Length | 128 |
| Sequences | 6 |
| Columns | 146 |
| Reading direction | forward |
| Mean pairwise identity | 83.61 |
| Shannon entropy | 0.29612 |
| G+C content | 0.29850 |
| Mean single sequence MFE | -25.95 |
| Consensus MFE | -17.43 |
| Energy contribution | -17.80 |
| Covariance contribution | 0.37 |
| Combinations/Pair | 1.15 |
| Mean z-score | -1.66 |
| Structure conservation index | 0.67 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.07 |
| SVM RNA-class probability | 0.529558 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 14267909 128 + 24543557 UAGAGAGAAAUAUUUAUUAUUUGCUUAUAAG-GCCAAACUAAAUAUUUGCCGCCGGCAAAUAUUUAAAA-GGUGUUAAGUGUUUCGUGGGAGUGAAGUUUUUAAACAG-GGAA---AAUAAAUCAAAUAAGUAU------------ .........(((((((((....(((((..(.-(((....((((((((((((...))))))))))))...-))).)))))).((((.((.(((.......)))...)).-))))---.........)))))))))------------ ( -25.20, z-score = -1.44, R) >droSim1.chr3L 13619744 128 + 22553184 UAGAGAGAAAUAUUUAUUAUUUGCUUAUAAG-GCCAAACUAAAUAUUUGCCGCCGGCAAAUGUUUAAAA-GGUGUUAAGUGUUUCGUGGGAGUGAAGUUUUUAAACAG-GAAA---AAUAAAUCAAAUAAAUAU------------ .........(((((((((....(((((..(.-(((....((((((((((((...))))))))))))...-))).)))))).((((.((.(((.......)))...)).-))))---.........)))))))))------------ ( -25.70, z-score = -1.77, R) >droSec1.super_0 6410695 128 + 21120651 UAGAGAGAAAUAUUUAUUAUUUGCUUAUAAG-GCCAAACUAAAUAUUUGCCGCCGGCAAAUGUUUAAAA-GGUGUUAAGUGUUUCGUGGGAGUGAAGUUUUUAAACAG-GAAA---AAUAAAUCAAAUAAAUAU------------ .........(((((((((....(((((..(.-(((....((((((((((((...))))))))))))...-))).)))))).((((.((.(((.......)))...)).-))))---.........)))))))))------------ ( -25.70, z-score = -1.77, R) >droYak2.chr3L 14361736 130 + 24197627 UAGAGAUAAAUAUUUAUUAUUUGCUUAUAAGCACAAAACUAAAUAUUUGCCGCCGGCAAAUGUUUAAAA-GGUGUUAAGUGAUUCGUGGGAGUGAAGUUUUUAAACAGCAAAA---AAGAAAUCAAAUAAAUAU------------ .........(((((((((.((((((....(((((.....((((((((((((...))))))))))))...-.)))))...(.((((....)))).)...........)))))).---..(....).)))))))))------------ ( -26.20, z-score = -2.16, R) >droEre2.scaffold_4784 14254723 128 + 25762168 UAGAGAAAAAUAUUUAUUAUUUGCUCAAAAG-CCGAAACUAAAUAUUUGCCGACGGCAAAUGUUUAAAA-GGUGUUAAGUGUUUCGUGGGAGUGAAGUUUUUAAACAG-GGAA---AAUAAAUCAAAUAAAUAU------------ .........(((((((((.((..(((....(-(.(((((((((((((((((...))))))))))))...-..........)))))))..)))..))((((((......-.)))---)))......)))))))))------------ ( -26.61, z-score = -2.49, R) >droAna3.scaffold_13337 18503392 140 - 23293914 CAAGCUUAAAUAUUUAUUGUUUGUUCAUAAA------GUUAAAUAUUUGCCGCCGGCAAAUGCCUAAAAAGGUGUCAAGUGUGUCGUGGGAGUGCGAAGGUUGAAUAUUGAGAUCAAAUAAGUGAAAUAAAUAACGAAAAGUUGCC ...((.....(((((.(((((((.((.(((.------(((..(((((((.(((((((....)))......)))).))))))).((((......))))......))).))).)).)))))))...)))))...(((.....))))). ( -26.30, z-score = -0.33, R) >consensus UAGAGAGAAAUAUUUAUUAUUUGCUUAUAAG_GCCAAACUAAAUAUUUGCCGCCGGCAAAUGUUUAAAA_GGUGUUAAGUGUUUCGUGGGAGUGAAGUUUUUAAACAG_GAAA___AAUAAAUCAAAUAAAUAU____________ .........(((((((((.....................((((((((((((...)))))))))))).....((.(((((..(((((......)))))..)))))))...................)))))))))............ (-17.43 = -17.80 + 0.37)
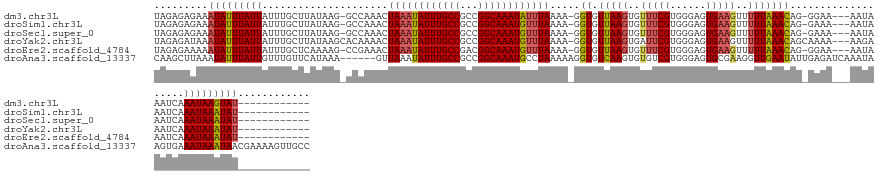

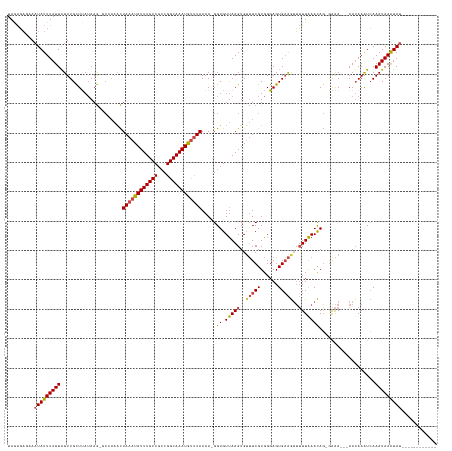
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:25:28 2011