
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 14,250,240 – 14,250,335 |
| Length | 95 |
| Max. P | 0.761653 |

| Location | 14,250,240 – 14,250,335 |
|---|---|
| Length | 95 |
| Sequences | 10 |
| Columns | 113 |
| Reading direction | forward |
| Mean pairwise identity | 84.29 |
| Shannon entropy | 0.31094 |
| G+C content | 0.37943 |
| Mean single sequence MFE | -23.70 |
| Consensus MFE | -16.83 |
| Energy contribution | -16.97 |
| Covariance contribution | 0.14 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.72 |
| Structure conservation index | 0.71 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.61 |
| SVM RNA-class probability | 0.761653 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 14250240 95 + 24543557 -------UUUUUUAAUAAAACAUGCAGCAAUAAAAUUCCAGCUGUGUUUACGUGAUUGGAAAACGAUUUUCCAUAGCGGCGAGCAUGAAAAAUUGUU----CCCGA------- -------..........((((..(((((............)))))))))..((...((((((.....))))))..))((.(((((........))))----)))..------- ( -20.90, z-score = -1.33, R) >droSim1.chr3L 13605114 95 + 22553184 ------CUUUUUUAAUAAAACACGCAGCAAUAAAAUUCCAGCUGUGUUUACGUGAUUGGAAAACGAUUUUCCAUAGCGGCGAGCAUGAAAAAUUGUU----CCCG-------- ------...............(((((((............)))))))....((...((((((.....))))))..))((.(((((........))))----))).-------- ( -23.20, z-score = -2.26, R) >droSec1.super_0 6397277 95 + 21120651 ------CUUUUUUAAUAAAACACGCAGCAAUAAAAUUCCAGCUGUGUUUACGUGAUUGGAAAACGAUUUUCCAUAGCGGCGAGCAUGAAAAAUUGUU----CCCG-------- ------...............(((((((............)))))))....((...((((((.....))))))..))((.(((((........))))----))).-------- ( -23.20, z-score = -2.26, R) >droYak2.chr3L 14348217 94 + 24197627 -------CUUUUUAAUAAAACACGCAGCAAUAAAAUUCCAGCUGUGUUUACGUGAUUGGAAAACGAUUUUCCAUAGCGGCGAGCAUGAAAAAUUGUU----CCCA-------- -------..............(((((((............)))))))....((...((((((.....))))))..))((.(((((........))))----))).-------- ( -23.20, z-score = -2.27, R) >droAna3.scaffold_13337 18491900 103 - 23293914 UUCAUUUUUUUUUAAUAAAACACGCAGCAAUAAAAUUCCAGCUGUGUUUACGUGAUUGGAAAACGAUUUUCCAUAGCGGCAGGCAUGAAAAAUUUUU----CUCAAA------ .......(((((((.......(((((((............)))))))...(((...((((((.....))))))..))).......))))))).....----......------ ( -19.70, z-score = -1.05, R) >dp4.chrXR_group6 6939058 107 + 13314419 ------GGUUUUUAAUAAAACACGCAGCAAUAAAAUUCCAGCUGUGUUUACGUGAUUGGAAAACGAUUUUCCAUAGCGGCGAGCAUGAAAAAUUGUUUGGCUUCAGCUACCAA ------.(((((....)))))(((((((............)))))))........((((.((((((((((.(((.((.....))))).))))))))))(((....))).)))) ( -26.70, z-score = -1.49, R) >droPer1.super_41 297045 107 + 728894 ------GGUUUUUAAUAAAACACGCAGCAAUAAAAUUCCAGCUGUGUUUACGUGAUUGGAAAACGAUUUUCCAUAGCGGCGAGCAUGAAAAAUUGUUUGGCUUCAGCUACCAA ------.(((((....)))))(((((((............)))))))........((((.((((((((((.(((.((.....))))).))))))))))(((....))).)))) ( -26.70, z-score = -1.49, R) >droWil1.scaffold_180949 3540707 84 + 6375548 -------GGUUUUAAUAAAACAAGCAGCAAUAAAAUUCCAGCUGUGUUUACGUGAUUGGAAAACGAUUUUCCAGUGCAUAAGGGGUGAGGG---------------------- -------.((((.....))))..(((((............))))).((((((((((((((((.....)))))))).))).....)))))..---------------------- ( -15.30, z-score = 0.12, R) >droVir3.scaffold_13049 3144820 98 - 25233164 ------UUUUUUUAUGAAAACACGCAGCAAUAAAAUUCCAGCUGUGUUUACGUGAUUGGAAAACGAUUUUCCAUGGCGGCGAGCAUGAAAAAUUGCU-GGCCCCG-------- ------.....(((((.((((..(((((............))))))))).))))).((((((.....)))))).((.(((.((((........))))-.))))).-------- ( -29.80, z-score = -2.67, R) >droMoj3.scaffold_6680 8089266 98 - 24764193 ------UUUUUUAAUAAAAACACGCAGCAAUAAAAUUCCAGCUGUGUUUACGUGAUUGGAAAACGAUUUUCCAUGGCGGCGAGCAUGAAAAAUUGCU-GGCCCCG-------- ------...............(((((((............))))))).........((((((.....)))))).((.(((.((((........))))-.))))).-------- ( -28.30, z-score = -2.46, R) >consensus _______UUUUUUAAUAAAACACGCAGCAAUAAAAUUCCAGCUGUGUUUACGUGAUUGGAAAACGAUUUUCCAUAGCGGCGAGCAUGAAAAAUUGUU____CCCG________ ........((((((.......(((((((............)))))))...(((...((((((.....))))))..))).......))))))...................... (-16.83 = -16.97 + 0.14)


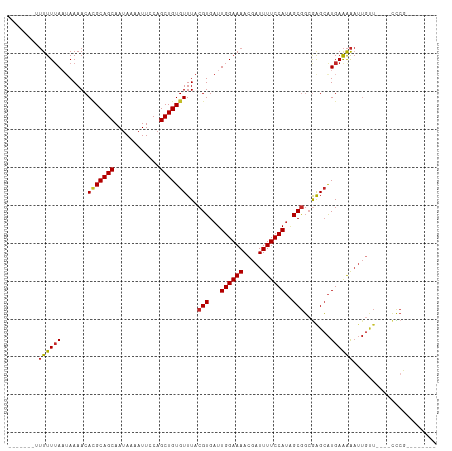
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:25:25 2011