| Sequence ID | dm3.chr3L |
|---|---|
| Location | 14,246,843 – 14,246,915 |
| Length | 72 |
| Max. P | 0.914033 |

| Location | 14,246,843 – 14,246,915 |
|---|---|
| Length | 72 |
| Sequences | 6 |
| Columns | 72 |
| Reading direction | forward |
| Mean pairwise identity | 84.63 |
| Shannon entropy | 0.30014 |
| G+C content | 0.46296 |
| Mean single sequence MFE | -20.55 |
| Consensus MFE | -17.11 |
| Energy contribution | -17.53 |
| Covariance contribution | 0.42 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.69 |
| Structure conservation index | 0.83 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.23 |
| SVM RNA-class probability | 0.914033 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
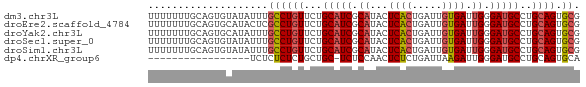
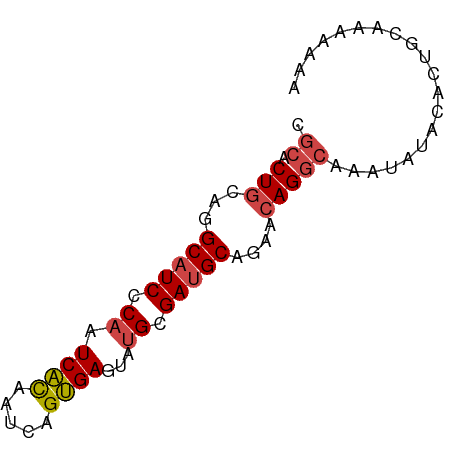
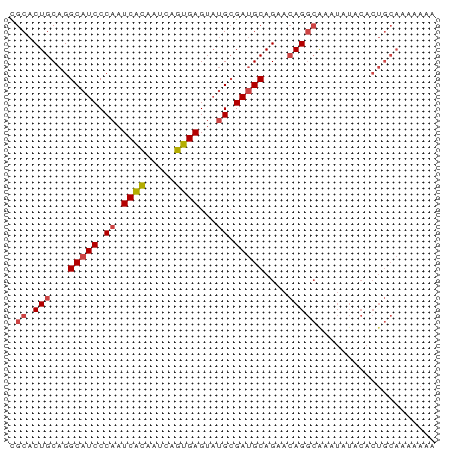
>dm3.chr3L 14246843 72 + 24543557 UUUUUUUGCAGUGUAUAUUUGCCUGUUCUGCAUCGCAUACUCACUGAUUGUGAUUGGGAUGCCUGCAGUGCG ....................((((((...(((((.((...((((.....)))).)).)))))..)))).)). ( -21.50, z-score = -1.84, R) >droEre2.scaffold_4784 14238256 72 + 25762168 UUUUUUUGCAGUGCAUACUCGCCUGUUCUGCAUCGCAUACUCACUGAUUGUGAUUGGGAUGCCUGCAGUGCG .......(((.((((.((......))...(((((.((...((((.....)))).)).))))).)))).))). ( -22.70, z-score = -1.71, R) >droYak2.chr3L 14344720 72 + 24197627 UUUUUUUGCAGUGCAUAUUUGCCUGUUCUGCAUCGCAUACUCACUGAUUGUGAUUGGGAUGCCUGCAGUGCG .......(((..(((....)))((((...(((((.((...((((.....)))).)).)))))..))))))). ( -22.70, z-score = -1.82, R) >droSec1.super_0 6393920 72 + 21120651 UUUUUUUGCAGUGUAUAUUUGCCUGUUCUGCAUCGCAUACUCACUGAUUGUGAUUGGGAUGCCUGCAGUGCG ....................((((((...(((((.((...((((.....)))).)).)))))..)))).)). ( -21.50, z-score = -1.84, R) >droSim1.chr3L 13601745 72 + 22553184 UUUUUUUGCAGUGUAUAUUUGCCUGUUCUGCAUCGCAUACUCACUGAUUGUGAUUGGGAUGCCUGCAGUGCG ....................((((((...(((((.((...((((.....)))).)).)))))..)))).)). ( -21.50, z-score = -1.84, R) >dp4.chrXR_group6 6935948 54 + 13314419 -----------------UCUCUCUCUGCUGC-UCUCCAACUCUCUGAUUAAGAUUGGGAUGCCUGCAGUGCA -----------------.....(.((((.((-..(((((.(((.......))))))))..))..)))).).. ( -13.40, z-score = -1.10, R) >consensus UUUUUUUGCAGUGUAUAUUUGCCUGUUCUGCAUCGCAUACUCACUGAUUGUGAUUGGGAUGCCUGCAGUGCG ....................((((((...(((((.((...((((.....)))).)).)))))..)))).)). (-17.11 = -17.53 + 0.42)
| Location | 14,246,843 – 14,246,915 |
|---|---|
| Length | 72 |
| Sequences | 6 |
| Columns | 72 |
| Reading direction | reverse |
| Mean pairwise identity | 84.63 |
| Shannon entropy | 0.30014 |
| G+C content | 0.46296 |
| Mean single sequence MFE | -18.30 |
| Consensus MFE | -14.23 |
| Energy contribution | -14.65 |
| Covariance contribution | 0.42 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.86 |
| Structure conservation index | 0.78 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.23 |
| SVM RNA-class probability | 0.912583 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
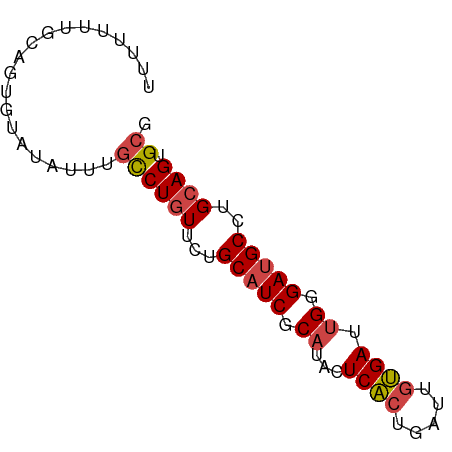
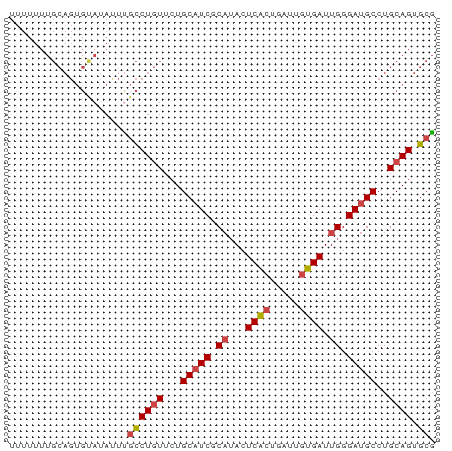
>dm3.chr3L 14246843 72 - 24543557 CGCACUGCAGGCAUCCCAAUCACAAUCAGUGAGUAUGCGAUGCAGAACAGGCAAAUAUACACUGCAAAAAAA .((.(((...(((((.((.((((.....))))...)).)))))....))))).................... ( -17.70, z-score = -1.88, R) >droEre2.scaffold_4784 14238256 72 - 25762168 CGCACUGCAGGCAUCCCAAUCACAAUCAGUGAGUAUGCGAUGCAGAACAGGCGAGUAUGCACUGCAAAAAAA .(((.((((.(((((.((.((((.....))))...)).)))))...((......)).)))).)))....... ( -20.10, z-score = -1.34, R) >droYak2.chr3L 14344720 72 - 24197627 CGCACUGCAGGCAUCCCAAUCACAAUCAGUGAGUAUGCGAUGCAGAACAGGCAAAUAUGCACUGCAAAAAAA .....((((((((((.((.((((.....))))...)).))))).......(((....))).)))))...... ( -21.10, z-score = -2.35, R) >droSec1.super_0 6393920 72 - 21120651 CGCACUGCAGGCAUCCCAAUCACAAUCAGUGAGUAUGCGAUGCAGAACAGGCAAAUAUACACUGCAAAAAAA .((.(((...(((((.((.((((.....))))...)).)))))....))))).................... ( -17.70, z-score = -1.88, R) >droSim1.chr3L 13601745 72 - 22553184 CGCACUGCAGGCAUCCCAAUCACAAUCAGUGAGUAUGCGAUGCAGAACAGGCAAAUAUACACUGCAAAAAAA .((.(((...(((((.((.((((.....))))...)).)))))....))))).................... ( -17.70, z-score = -1.88, R) >dp4.chrXR_group6 6935948 54 - 13314419 UGCACUGCAGGCAUCCCAAUCUUAAUCAGAGAGUUGGAGA-GCAGCAGAGAGAGA----------------- ....((((..((.((((((((((.....)))).)))).))-)).)))).......----------------- ( -15.50, z-score = -1.82, R) >consensus CGCACUGCAGGCAUCCCAAUCACAAUCAGUGAGUAUGCGAUGCAGAACAGGCAAAUAUACACUGCAAAAAAA .((.(((...(((((.((.((((.....))))...)).)))))....))))).................... (-14.23 = -14.65 + 0.42)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:25:23 2011