
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 13,995,198 – 13,995,270 |
| Length | 72 |
| Max. P | 0.971907 |

| Location | 13,995,198 – 13,995,270 |
|---|---|
| Length | 72 |
| Sequences | 6 |
| Columns | 72 |
| Reading direction | forward |
| Mean pairwise identity | 77.96 |
| Shannon entropy | 0.42580 |
| G+C content | 0.45449 |
| Mean single sequence MFE | -22.18 |
| Consensus MFE | -11.40 |
| Energy contribution | -12.85 |
| Covariance contribution | 1.45 |
| Combinations/Pair | 1.19 |
| Mean z-score | -2.55 |
| Structure conservation index | 0.51 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.86 |
| SVM RNA-class probability | 0.971907 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
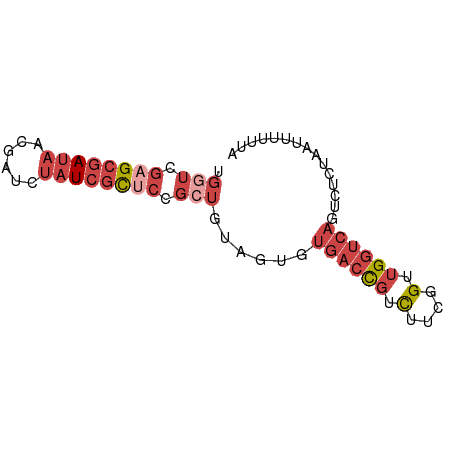
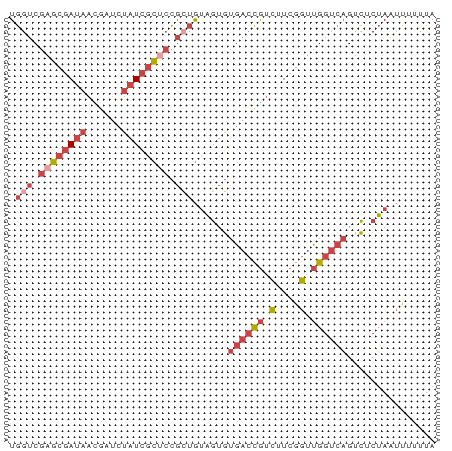
>dm3.chr3L 13995198 72 + 24543557 UGGUCGAGCGAUAACGAUCUAUCGCUCCGCUGUAGUGUGACCGUCUUCGGUUGGUCAGUCUCUAAUUUUUUA .(((.((((((((......)))))))).))).(((.(((((((.(....).))))))..).)))........ ( -24.50, z-score = -3.24, R) >droWil1.scaffold_180698 5143054 71 - 11422946 UAGACGAG-AACAGUAUCCAAUACUUAUCGCAACUUUUGAGUAAUGUAAGACGGAAUGUAUCGAUAUUGUUA ........-(((((((((((.((((((..........)))))).)).......((.....))))))))))). ( -11.70, z-score = -0.32, R) >droSim1.chr3L 13350873 72 + 22553184 UGGUCGAGCGAUAACGAUGUAUCGCUCCGCUGUAGUGUGACCGUCUUCGGUUGGUCAGUCUCUAAUUUUUUA .(((.((((((((......)))))))).))).(((.(((((((.(....).))))))..).)))........ ( -24.30, z-score = -3.07, R) >droSec1.super_0 6141836 72 + 21120651 UGGUCGAGCGAUAACGAUCUAUCGCUCCGCUGUAGUGUGACCGUCUUCGGUUGGUCAGUCUCUAAUUUUUUA .(((.((((((((......)))))))).))).(((.(((((((.(....).))))))..).)))........ ( -24.50, z-score = -3.24, R) >droYak2.chr3L 14082883 72 + 24197627 UGGUCGAGCGAUAACGAUCUAUCGCCCCGAUGUAGUGUGACCGUCUUCGGUUGGUCAGUCGCUAAUUUUUUA ..((((.((((((......))))))..)))).(((((((((((.(....).))))))..)))))........ ( -24.80, z-score = -2.81, R) >droEre2.scaffold_4784 13987201 72 + 25762168 UGGUCGAGCGAUAAUGAUCUAUCGCCCCGCUGUAGUGUGACCGUCUUCGGUUGGUCAGUCGCUAAUUUUUUA .(((.(.((((((......)))))).).))).(((((((((((.(....).))))))..)))))........ ( -23.30, z-score = -2.63, R) >consensus UGGUCGAGCGAUAACGAUCUAUCGCUCCGCUGUAGUGUGACCGUCUUCGGUUGGUCAGUCUCUAAUUUUUUA ((..(((((((((......))))))))...........(((((....))))))..))............... (-11.40 = -12.85 + 1.45)

| Location | 13,995,198 – 13,995,270 |
|---|---|
| Length | 72 |
| Sequences | 6 |
| Columns | 72 |
| Reading direction | reverse |
| Mean pairwise identity | 77.96 |
| Shannon entropy | 0.42580 |
| G+C content | 0.45449 |
| Mean single sequence MFE | -15.82 |
| Consensus MFE | -11.31 |
| Energy contribution | -12.07 |
| Covariance contribution | 0.75 |
| Combinations/Pair | 1.29 |
| Mean z-score | -1.70 |
| Structure conservation index | 0.72 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.60 |
| SVM RNA-class probability | 0.953996 |
| Prediction | RNA |
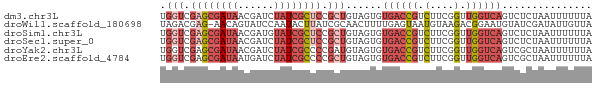
Download alignment: ClustalW | MAF
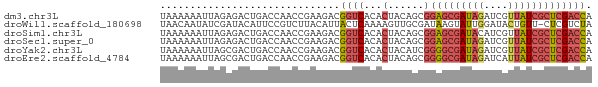
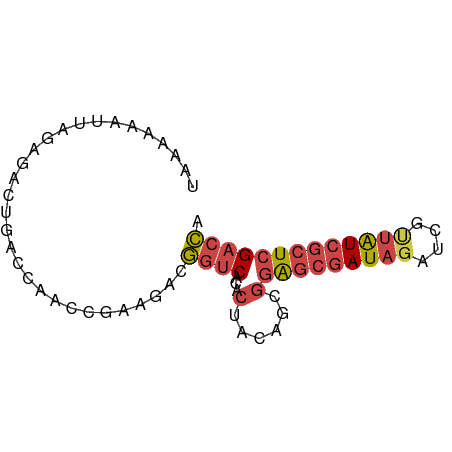
>dm3.chr3L 13995198 72 - 24543557 UAAAAAAUUAGAGACUGACCAACCGAAGACGGUCACACUACAGCGGAGCGAUAGAUCGUUAUCGCUCGACCA ..............................((((.(.(....).)(((((((((....))))))))))))). ( -17.70, z-score = -2.36, R) >droWil1.scaffold_180698 5143054 71 + 11422946 UAACAAUAUCGAUACAUUCCGUCUUACAUUACUCAAAAGUUGCGAUAAGUAUUGGAUACUGUU-CUCGUCUA .((((.(((((((((..((((.(((...........))).)).))...))))).)))).))))-........ ( -8.40, z-score = 0.17, R) >droSim1.chr3L 13350873 72 - 22553184 UAAAAAAUUAGAGACUGACCAACCGAAGACGGUCACACUACAGCGGAGCGAUACAUCGUUAUCGCUCGACCA ..............................((((.(.(....).)((((((((......)))))))))))). ( -17.20, z-score = -2.47, R) >droSec1.super_0 6141836 72 - 21120651 UAAAAAAUUAGAGACUGACCAACCGAAGACGGUCACACUACAGCGGAGCGAUAGAUCGUUAUCGCUCGACCA ..............................((((.(.(....).)(((((((((....))))))))))))). ( -17.70, z-score = -2.36, R) >droYak2.chr3L 14082883 72 - 24197627 UAAAAAAUUAGCGACUGACCAACCGAAGACGGUCACACUACAUCGGGGCGAUAGAUCGUUAUCGCUCGACCA ........(((.(..(((((..........)))))).))).....(((((((((....)))))))))..... ( -16.90, z-score = -1.26, R) >droEre2.scaffold_4784 13987201 72 - 25762168 UAAAAAAUUAGCGACUGACCAACCGAAGACGGUCACACUACAGCGGGGCGAUAGAUCAUUAUCGCUCGACCA ...........((.(((....((((....)))).......)))))(((((((((....)))))))))..... ( -17.00, z-score = -1.91, R) >consensus UAAAAAAUUAGAGACUGACCAACCGAAGACGGUCACACUACAGCGGAGCGAUAGAUCGUUAUCGCUCGACCA ..............................((((...(......)(((((((((....))))))))))))). (-11.31 = -12.07 + 0.75)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:24:56 2011