
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 5,202,385 – 5,202,519 |
| Length | 134 |
| Max. P | 0.725194 |

| Location | 5,202,385 – 5,202,519 |
|---|---|
| Length | 134 |
| Sequences | 12 |
| Columns | 136 |
| Reading direction | forward |
| Mean pairwise identity | 76.38 |
| Shannon entropy | 0.51463 |
| G+C content | 0.38546 |
| Mean single sequence MFE | -30.49 |
| Consensus MFE | -18.30 |
| Energy contribution | -17.12 |
| Covariance contribution | -1.17 |
| Combinations/Pair | 1.38 |
| Mean z-score | -1.04 |
| Structure conservation index | 0.60 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.51 |
| SVM RNA-class probability | 0.725194 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 5202385 134 + 23011544 GAGGCAAAGUGGCAUAUUUGGGAAUAUACCUUCCCAAUUGUGUCUCAAAUACUCCAAAAACAACUAUGUAAU-AGGGACGAACGGCAUCCUGAAUAGUGC-AUCAUACUAUAGCAGAUGACUUGUGUUCGUGCACA (((.......((((((.(((((((......))))))).)))))).......))).........((((...))-))(.((((((((((((.((.((((((.-....))))))..)))))).))...)))))).)... ( -36.04, z-score = -1.60, R) >droSim1.chr2L 4999630 134 + 22036055 GAGUCAAAGUGGCAUAUUUGGGAAUGUACCUUCCCAAUUGUGUCUCAAAUACUCCAAAAAUAACUAUGUAAU-AGGGGCGAACGGCAUCCUGAAUAGUGC-AUCAUACUAUAGCAGAUGACUUGUGUUCGUGCACA ((((((..(.((((((.(((((((......))))))).)))))).)....................(((..(-((((((.....)).))))).((((((.-....)))))).)))..))))))((((....)))). ( -37.30, z-score = -1.71, R) >droSec1.super_5 3283596 134 + 5866729 GAGUCAAAGUGGCAUAUUUGGGAAUAUACCUUCCCAAUUGUGUCUCAAAUACUCCAAAAUCAACUAUGUAAU-AGGGACGAACGGCAUCCUGAAUAGUGC-AUCAUACUAUAGCAGAUGACUUGUGUUCGUCCACA ((((....(.((((((.(((((((......))))))).)))))).)....)))).........((((...))-))((((((((((((((.((.((((((.-....))))))..)))))).))...))))))))... ( -41.30, z-score = -3.59, R) >droYak2.chr2L 11999171 134 - 22324452 GAGUCAAAGUGGCAUAUUUGAGAAUAGCCCUUCCCAAUUGUGUCUCAAAUACUCCGAAAACAACUAUGCAAU-AGGGACGAACGGCAUCCUGAAUAGUGU-AUCAUAUUAUAACAGAUGACUUGUGUUCGUCCACA ((((....(.((((((.(((.(((......))).))).)))))).)....)))).........((((...))-))((((((((((((((.((.(((((((-...)))))))..)))))).))...))))))))... ( -32.00, z-score = -1.31, R) >droEre2.scaffold_4929 5280404 134 + 26641161 GAGUCAAAGUGGCAUAUUUGGGUAUAUCCCUGCCCAUUUGUGUCUCAAAUACUCCCAUAACAACUUUGUAGU-AGGGAAGAACGGCAUCCUGAAUAGUGC-AUUAUAUUAUAGCAGAUGACUUGAGUUCGUCCACA ((((....(.((((((..((((((......))))))..)))))).)....))))......(.(((....)))-.)(((.((((((((((.((.((((((.-....))))))..)))))).))...)))).)))... ( -33.30, z-score = -0.73, R) >droAna3.scaffold_12916 6217480 128 - 16180835 GAGCCAAAGUGGCAUAUAUCAGAAUCUCUAAUCCGAUUUGUGUCUCAAAUGCUCCCAAACUAA-------GC-AAAGAUGAACGGCAUCCUGUAUAGUUCCAUCAUAUUAUAACAGAUGACUUGUGUUCAACUACA ((((....(.((((((.(((.((........)).))).)))))).)....)))).........-------..-..((.(((((((((((.((((((((........))))).))))))).))...))))).))... ( -26.30, z-score = -1.16, R) >dp4.chr4_group3 9001931 134 - 11692001 GAGUGAAAGUGGCAUAUUGUAUAACCAUUAUUUGC-UCUGUGUCUCAACUGCUCCCAAAGCAACUUCUUUUCGAGAGACAAACGGCAUCCUGUCUAGUGG-AUCAUACUAAAACAGAUGACUUGUGUUUGUCUACA (((...((((((((((..(((...........)))-..)))))).....((((.....))))))))...)))...((((((((((((((.(((.(((((.-....)))))..))))))).))...))))))))... ( -30.20, z-score = -0.25, R) >droPer1.super_1 6101976 134 - 10282868 GAGUGAAAGUGGCAUAUUGUAUAACCAUUAUUUGC-UCUGUGUCUCAACUGCUCCCAAAGCAACUUCUUUUCGAGAGACAAACGGCAUCCUGUCUAGUGG-AUCAUACUAAAACAGAUGACUUGUGUUUGUCUACA (((...((((((((((..(((...........)))-..)))))).....((((.....))))))))...)))...((((((((((((((.(((.(((((.-....)))))..))))))).))...))))))))... ( -30.20, z-score = -0.25, R) >droWil1.scaffold_180772 7827562 131 - 8906247 GAGUUAAAGUGGCAUAUUUAAGUUGUAUUUUCCUCAUUUGUGUCUCAAAUACUCCAAAAGAAAA----CUUU-CAAGACGAACGGCAUCCUGCAUAAAUUAUUAAUUUUAUAACAGAUGACUUGUGUUCGUGUGCA ((((....(.((((((..(.((..(....)..)).)..)))))).)....))))..........----....-..(.((((((((((((.((.(((((........)))))..)))))).))...)))))).)... ( -20.90, z-score = 0.63, R) >droVir3.scaffold_12963 2584658 126 - 20206255 GAGUCAAAGUGGCAUAUUUUAGGAA-UUUAACCGAAUUUGUGUCUCAAAUGCUCCAAAACUAA------CUA-AAAGACGAACGGCAUCCUGUUUAA-UCCCUCAUAUUAAAACAGAUGACUUGUGUUCGUGUAC- ((((....(.((((((..((.((..-.....)).))..)))))).)....)))).........------...-..(.((((((((((((.(((((..-............))))))))).))...)))))).)..- ( -23.94, z-score = -0.71, R) >droMoj3.scaffold_6500 9689630 128 - 32352404 GAGUCAAAGUGGCAUAUUUUGAAAUAUAAAUUACGAUUUGUGUCUCAAAUGCUCCAAAAUCAA------UUU-AUAGACGAACGGCAUCCUGUUUAA-UUAUUCAUAUUACAACAGAUGACUUGUGUUCGUGUACA ........(..((((...((((.((((((((....)))))))).))))))))..)........------...-.((.((((((((((((.(((((((-(.......)))).)))))))).))...)))))).)).. ( -26.70, z-score = -1.20, R) >droGri2.scaffold_15252 2147324 127 - 17193109 GAGUCAAAGUGGCAUAUUUUGGAGU-UCUCUUCCGAUUUGUGUCUCAGAAGCUCCAAAAUCAA------CUU-AAAUGCGAACGGCAUCCUGUUUAA-UCCCUUAUAUUAUAACAGAUGACUUGUGUUGGUGUACA ((((((((((.....((((((((((-(.(((...(((....)))..))))))))))))))..)------)))-..((((.....)))).((((((((-(.......)))).))))).))))))............. ( -27.70, z-score = -0.56, R) >consensus GAGUCAAAGUGGCAUAUUUGAGAAUAUAUCUUCCCAUUUGUGUCUCAAAUACUCCAAAAACAACU___UAUU_AGAGACGAACGGCAUCCUGUAUAGUGC_AUCAUAUUAUAACAGAUGACUUGUGUUCGUCUACA ((((....(.((((((..((..............))..)))))).)....)))).....................(.((((((((((((.((..((((........))))...)))))).))...)))))).)... (-18.30 = -17.12 + -1.17)

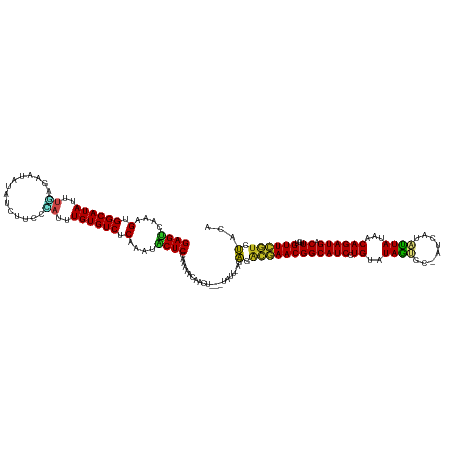

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:18:18 2011