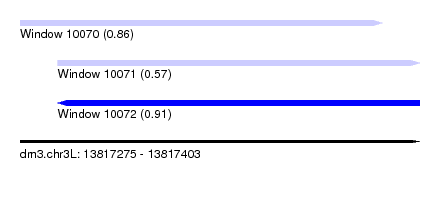

| Sequence ID | dm3.chr3L |
|---|---|
| Location | 13,817,275 – 13,817,403 |
| Length | 128 |
| Max. P | 0.907250 |

| Location | 13,817,275 – 13,817,391 |
|---|---|
| Length | 116 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 94.39 |
| Shannon entropy | 0.09833 |
| G+C content | 0.40230 |
| Mean single sequence MFE | -32.96 |
| Consensus MFE | -28.54 |
| Energy contribution | -29.74 |
| Covariance contribution | 1.20 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.03 |
| Structure conservation index | 0.87 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.95 |
| SVM RNA-class probability | 0.859138 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 13817275 116 + 24543557 UCCUUCUAGGCCAUAUAUUUUAGCUAAUCCCUGAGAUAUGCAGCCAAAAGCUCUUAAAAGUGCGGAGUACUUUUUAGAGCUUCUAGACUUUUCAAGGCUUACAU----CUGCUGUGGAUG ....(((((((..((((((((((.......))))))))))..)))..(((((((.((((((((...)))))))).))))))).))))...((((.(((......----..))).)))).. ( -35.60, z-score = -2.55, R) >droSim1.chr3L 13185814 116 + 22553184 UCCUUCUAUGCCAUAUAUUUUACCUAAUCUCUGAGAUAUGCAGCCCAAAGCUCUUAAAAGUGCGGAGUACUUUUUACAGCUCUUAGACUUUUCAAGGCUUACAU----CUGCUGUGGAUA ..........((((((((((((.........))))))))((((....((((.(((((((((..(((((..........)))))...)))))).)))))))....----)))).))))... ( -24.40, z-score = -0.48, R) >droSec1.super_0 5960645 116 + 21120651 UCCUUCUAGGCCAUAUAUUUUAGCUAAUCCCUGAGAUAUGCAGCCAAAAGCUCUUAAAAGUUCGGACUACUUUUUAGAGCUCUUAGACUUUUCAAGGCUUACAU----CUGCUGUGGAUG ....(((((((..((((((((((.......))))))))))..)))...((((((.((((((.......)))))).))))))..))))...((((.(((......----..))).)))).. ( -31.90, z-score = -2.14, R) >droYak2.chr3L 13899474 120 + 24197627 UCCUUGUAGGCCAUAUAUUUUAGCUAAUCCCUGAGAUAUGCAGCCAAAAGCUCUUAAAAGUGCGGAGUACUUUUUAGAGCUCCUAGACUUUUCAGGGCUUACAUACAUCUGCUGUGGAUG (((..((((((..((((((((((.......))))))))))..)))..(((((((.((((((..(((((.((....)).)))))...)))))).))))))).........)))...))).. ( -38.10, z-score = -2.69, R) >droEre2.scaffold_4784 13805839 116 + 25762168 UCCUUCUAGGCCAUAUAUUUUAGCUAAUCCCUGAGAUAUGCAGCCAAAAGCUCUUAAAAGUGCGGAGUACUUUUUAGAGCUCUUAGACUUUUCAAGGCUUACAU----CUGCUGUGGAUG ....(((((((..((((((((((.......))))))))))..)))...((((((.((((((((...)))))))).))))))..))))...((((.(((......----..))).)))).. ( -34.80, z-score = -2.30, R) >consensus UCCUUCUAGGCCAUAUAUUUUAGCUAAUCCCUGAGAUAUGCAGCCAAAAGCUCUUAAAAGUGCGGAGUACUUUUUAGAGCUCUUAGACUUUUCAAGGCUUACAU____CUGCUGUGGAUG ....(((((((..((((((((((.......))))))))))..)))...((((((.((((((((...)))))))).))))))..))))...((((.(((............))).)))).. (-28.54 = -29.74 + 1.20)

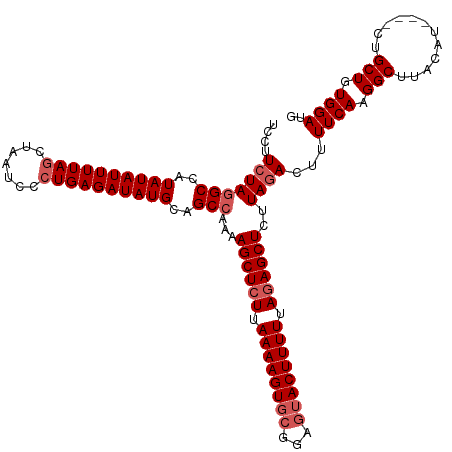
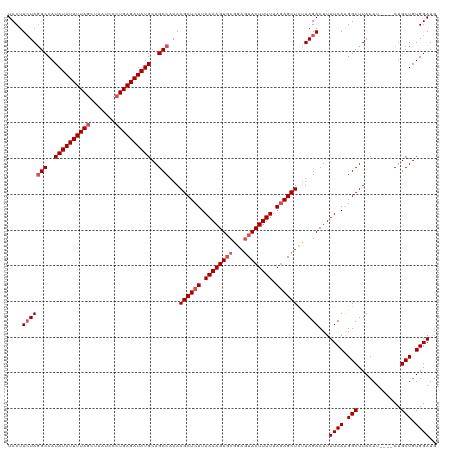
| Location | 13,817,287 – 13,817,403 |
|---|---|
| Length | 116 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 94.73 |
| Shannon entropy | 0.09232 |
| G+C content | 0.39713 |
| Mean single sequence MFE | -31.86 |
| Consensus MFE | -28.66 |
| Energy contribution | -29.46 |
| Covariance contribution | 0.80 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.40 |
| Structure conservation index | 0.90 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.15 |
| SVM RNA-class probability | 0.567379 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 13817287 116 + 24543557 AUAUAUUUUAGCUAAUCCCUGAGAUAUGCAGCCAAAAGCUCUUAAAAGUGCGGAGUACUUUUUAGAGCUUCUAGACUUUUCAAGGCUUACAU----CUGCUGUGGAUGGGUUCCGCUUAA .((((((((((.......)))))))))).((((..(((((((.((((((((...)))))))).)))))))...((....))..)))).....----.....(((((.....))))).... ( -33.00, z-score = -1.58, R) >droSim1.chr3L 13185826 116 + 22553184 AUAUAUUUUACCUAAUCUCUGAGAUAUGCAGCCCAAAGCUCUUAAAAGUGCGGAGUACUUUUUACAGCUCUUAGACUUUUCAAGGCUUACAU----CUGCUGUGGAUAGGUUCCGCUUAA ..((((((((.........))))))))((((....((((.(((((((((..(((((..........)))))...)))))).)))))))....----)))).(((((.....))))).... ( -26.80, z-score = -0.63, R) >droSec1.super_0 5960657 116 + 21120651 AUAUAUUUUAGCUAAUCCCUGAGAUAUGCAGCCAAAAGCUCUUAAAAGUUCGGACUACUUUUUAGAGCUCUUAGACUUUUCAAGGCUUACAU----CUGCUGUGGAUGGGUUCCGCUUAA .((((((((((.......)))))))))).((((...((((((.((((((.......)))))).))))))....((....))..)))).....----.....(((((.....))))).... ( -29.20, z-score = -1.07, R) >droYak2.chr3L 13899486 120 + 24197627 AUAUAUUUUAGCUAAUCCCUGAGAUAUGCAGCCAAAAGCUCUUAAAAGUGCGGAGUACUUUUUAGAGCUCCUAGACUUUUCAGGGCUUACAUACAUCUGCUGUGGAUGGGCUCCGCUUAA ..(((((((((.......)))))))))((((((..(((((((.((((((..(((((.((....)).)))))...)))))).))))))).....(((((.....)))))))))..)).... ( -38.20, z-score = -2.42, R) >droEre2.scaffold_4784 13805851 116 + 25762168 AUAUAUUUUAGCUAAUCCCUGAGAUAUGCAGCCAAAAGCUCUUAAAAGUGCGGAGUACUUUUUAGAGCUCUUAGACUUUUCAAGGCUUACAU----CUGCUGUGGAUGGGUUCCGCUUAA .((((((((((.......)))))))))).((((...((((((.((((((((...)))))))).))))))....((....))..)))).....----.....(((((.....))))).... ( -32.10, z-score = -1.31, R) >consensus AUAUAUUUUAGCUAAUCCCUGAGAUAUGCAGCCAAAAGCUCUUAAAAGUGCGGAGUACUUUUUAGAGCUCUUAGACUUUUCAAGGCUUACAU____CUGCUGUGGAUGGGUUCCGCUUAA .((((((((((.......)))))))))).((((...((((((.((((((((...)))))))).))))))....((....))..))))..............(((((.....))))).... (-28.66 = -29.46 + 0.80)

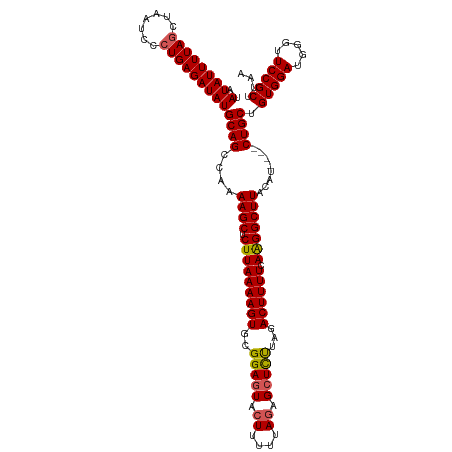
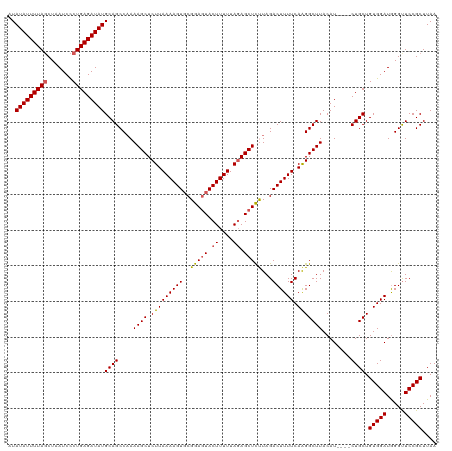
| Location | 13,817,287 – 13,817,403 |
|---|---|
| Length | 116 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 94.73 |
| Shannon entropy | 0.09232 |
| G+C content | 0.39713 |
| Mean single sequence MFE | -31.52 |
| Consensus MFE | -29.06 |
| Energy contribution | -29.30 |
| Covariance contribution | 0.24 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.90 |
| Structure conservation index | 0.92 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.19 |
| SVM RNA-class probability | 0.907250 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
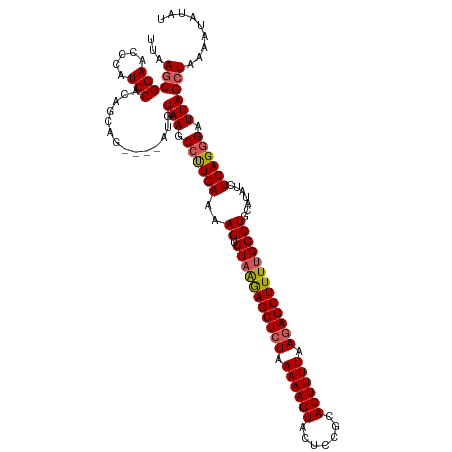
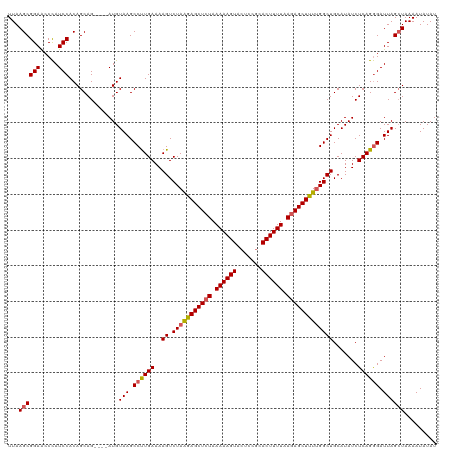
>dm3.chr3L 13817287 116 - 24543557 UUAAGCGGAACCCAUCCACAGCAG----AUGUAAGCCUUGAAAAGUCUAGAAGCUCUAAAAAGUACUCCGCACUUUUAAGAGCUUUUGGCUGCAUAUCUCAGGGAUUAGCUAAAAUAUAU ...((((((.....))).......----...(((.((((((..((.(((((((((((.((((((.......)))))).))))))))))))).......)))))).))))))......... ( -32.00, z-score = -2.32, R) >droSim1.chr3L 13185826 116 - 22553184 UUAAGCGGAACCUAUCCACAGCAG----AUGUAAGCCUUGAAAAGUCUAAGAGCUGUAAAAAGUACUCCGCACUUUUAAGAGCUUUGGGCUGCAUAUCUCAGAGAUUAGGUAAAAUAUAU .........(((((..(.(.(.((----((((.(((((...........((((((.(.((((((.......)))))).).)))))))))))..))))))).).)..)))))......... ( -26.20, z-score = -0.45, R) >droSec1.super_0 5960657 116 - 21120651 UUAAGCGGAACCCAUCCACAGCAG----AUGUAAGCCUUGAAAAGUCUAAGAGCUCUAAAAAGUAGUCCGAACUUUUAAGAGCUUUUGGCUGCAUAUCUCAGGGAUUAGCUAAAAUAUAU ...((((((.....))).......----...(((.((((((..((.(((((((((((.((((((.......)))))).))))))))))))).......)))))).))))))......... ( -32.20, z-score = -2.44, R) >droYak2.chr3L 13899486 120 - 24197627 UUAAGCGGAGCCCAUCCACAGCAGAUGUAUGUAAGCCCUGAAAAGUCUAGGAGCUCUAAAAAGUACUCCGCACUUUUAAGAGCUUUUGGCUGCAUAUCUCAGGGAUUAGCUAAAAUAUAU ...((((((.....)))...(((......)))...((((((..((.(((((((((((.((((((.......)))))).))))))))))))).......))))))....)))......... ( -35.20, z-score = -2.00, R) >droEre2.scaffold_4784 13805851 116 - 25762168 UUAAGCGGAACCCAUCCACAGCAG----AUGUAAGCCUUGAAAAGUCUAAGAGCUCUAAAAAGUACUCCGCACUUUUAAGAGCUUUUGGCUGCAUAUCUCAGGGAUUAGCUAAAAUAUAU ...((((((.....))).......----...(((.((((((..((.(((((((((((.((((((.......)))))).))))))))))))).......)))))).))))))......... ( -32.00, z-score = -2.32, R) >consensus UUAAGCGGAACCCAUCCACAGCAG____AUGUAAGCCUUGAAAAGUCUAAGAGCUCUAAAAAGUACUCCGCACUUUUAAGAGCUUUUGGCUGCAUAUCUCAGGGAUUAGCUAAAAUAUAU ...((((((.....)))...(((......)))...((((((..((.(((((((((((.((((((.......)))))).))))))))))))).......))))))....)))......... (-29.06 = -29.30 + 0.24)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:24:35 2011