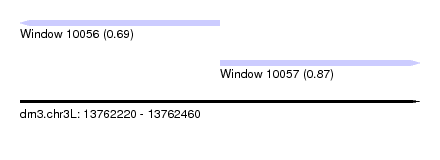
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 13,762,220 – 13,762,460 |
| Length | 240 |
| Max. P | 0.866127 |

| Location | 13,762,220 – 13,762,340 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 98.00 |
| Shannon entropy | 0.03610 |
| G+C content | 0.44833 |
| Mean single sequence MFE | -31.96 |
| Consensus MFE | -30.64 |
| Energy contribution | -30.32 |
| Covariance contribution | -0.32 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.31 |
| Structure conservation index | 0.96 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.43 |
| SVM RNA-class probability | 0.693896 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
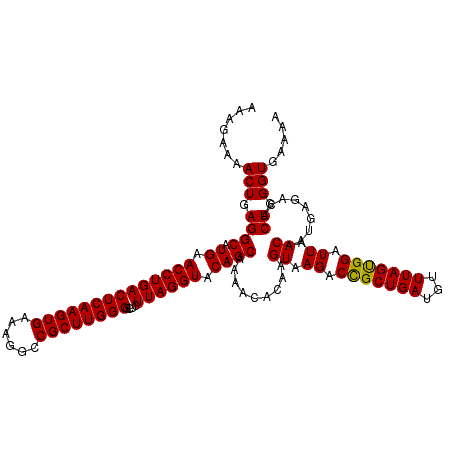
>dm3.chr3L 13762220 120 - 24543557 AAAGAAAACUGAGGCAUGAACCUGACUCAAGUGAAAGGCCGCUUGGGGCUCUUAGGUACAGCAAAACACAAGUAAGACCGCUGAUGUUUAGCGGAUUAACAAUGAGCCACUGGGUGAAAA .......(((.((((.((.((((((((((((((......))))))))....)))))).)))).........((.((.(((((((...))))))).)).)).........)).)))..... ( -32.60, z-score = -1.20, R) >droSim1.chr3L 13147815 120 - 22553184 AAAGAAAACUGAGGCAUGAACCUGACUCAAGUGAAAGGCCGCUUGGGGCUCUUAGGUACAGCAAAACACAAGUAAGACCGCUGAUGUUUAGUGGAUUAACAAUGAGACACUGGGUGAAAA .......(((.((((.((.((((((((((((((......))))))))....)))))).)))).........((.((.(((((((...))))))).)).)).........)).)))..... ( -30.70, z-score = -1.15, R) >droSec1.super_0 5923768 120 - 21120651 AAAGAAAACUGAGGCAUGAACCUGACUCAAGUGAAAGGCCGCUUGGGGCUCUUAGGUACAGCAAAACACAAGUAAGACUGCUGAUGUUUAGUGGAUUAACAAUGAGACACUGGGUGAAAA .......(((...((.((.((((((((((((((......))))))))....)))))).)))).(((((..((((....))))..)))))((((..(((....)))..)))).)))..... ( -29.80, z-score = -0.96, R) >droYak2.chr3L 13860114 120 - 24197627 AAAGAAAACUGAGGCAUGAACCUGACUCAAGUGAAAGGCCGCUUGGGGCUCUUAGGUACAGCCAAACACAAGUAAGACCGCUGAUGUUUAGUGGAUUAACAAGGAGACACUGGGUGAAAA ............(((.((.((((((((((((((......))))))))....)))))).)))))...(((.(((....(((((((...)))))))........(....))))..))).... ( -36.00, z-score = -2.15, R) >droEre2.scaffold_4784 13768519 120 - 25762168 AAAGAAAACUGAGGCAUGAACCUGACUCAAGUGAAAGGCCGCUUGGGGCUCUUAGGUACAGCAAAACACAAGUAAGACCGCUGAUGUUUAGUGGAUUAACAAUGAGACACUGGGUGAAAG .......(((.((((.((.((((((((((((((......))))))))....)))))).)))).........((.((.(((((((...))))))).)).)).........)).)))..... ( -30.70, z-score = -1.10, R) >consensus AAAGAAAACUGAGGCAUGAACCUGACUCAAGUGAAAGGCCGCUUGGGGCUCUUAGGUACAGCAAAACACAAGUAAGACCGCUGAUGUUUAGUGGAUUAACAAUGAGACACUGGGUGAAAA .......(((.((((.((.((((((((((((((......))))))))....)))))).)))).........((.((.(((((((...))))))).)).)).........)).)))..... (-30.64 = -30.32 + -0.32)


| Location | 13,762,340 – 13,762,460 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 97.99 |
| Shannon entropy | 0.03423 |
| G+C content | 0.41108 |
| Mean single sequence MFE | -26.78 |
| Consensus MFE | -25.34 |
| Energy contribution | -25.34 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.73 |
| Structure conservation index | 0.95 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.98 |
| SVM RNA-class probability | 0.866127 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
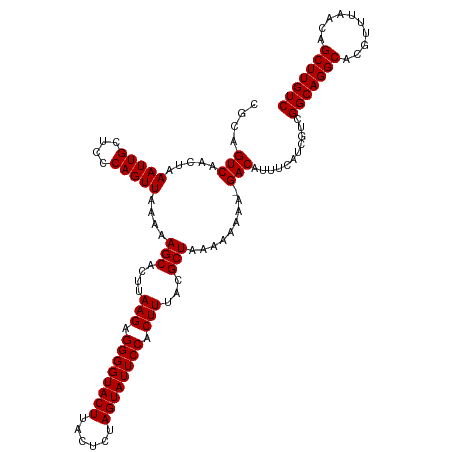
>dm3.chr3L 13762340 120 + 24543557 CGCAGUCAACUAAAUUGCUCCCAGUUAAAAAGCACUUAAGAGGGGUACUUACUCUAGUAUUCCACUUUACGCUAAAAAAAAAGACAUUUCAUCGUCGGCAGGCACGUUUAACAGCUUGUC .(((((.......)))))............(((....(((.((((((((......)))))))).)))...))).........(((........)))(((((((..........))))))) ( -26.80, z-score = -1.85, R) >droSim1.chr3L 13147935 119 + 22553184 UGCAGUCAACUAAAUUGCUCCCAGUUAAAAAGCACUUAAGAGGGGUACUUACUCUAGUAUUCCACUUUACGCUAAAAAAAA-GACAUUUCAUCGUCGGCAGGCACGUUUAACAGCUUGUC .(((((.......)))))............(((....(((.((((((((......)))))))).)))...)))........-(((........)))(((((((..........))))))) ( -26.50, z-score = -1.62, R) >droSec1.super_0 5923888 119 + 21120651 CGCAGUCAACUAAAUUGCUCCCAGUUAAAAAGCACUUAAGAGGGGUACUUACUCUAGUAUUCCACUUUACGCUAAAAAAAA-GACAUUUCAUCGUCGGCAGGCACGUUUAACAGCUUGUC .(((((.......)))))............(((....(((.((((((((......)))))))).)))...)))........-(((........)))(((((((..........))))))) ( -26.80, z-score = -1.90, R) >droYak2.chr3L 13860234 119 + 24197627 CUCAGUCGACUAAAUUGCUCCCAGUUAAAAAGCACUUAAGAGGGGUACUUACUCUAGUAUUCCACUUUACGCUAAAAAAAA-GACAUUUCAUCGUCGGCAGGCACGUUUAACAGCUUGUC ....((((((..(((((....)))))....(((....(((.((((((((......)))))))).)))...)))........-...........)))))).((((.((......)).)))) ( -26.90, z-score = -1.79, R) >droEre2.scaffold_4784 13768639 119 + 25762168 CUCAGUCGGCUAAAUUGCUCCCAGUUAAAAAGCACUUAAGAGGGGUACUUACUCUAGUAUUCCACUUUACGCUAAAAAAAA-GACAUUUCAUCGUCGGCAGGCACGUUUAACAGCUUGUC ....((((((......)))...........(((....(((.((((((((......)))))))).)))...)))........-)))...........(((((((..........))))))) ( -26.90, z-score = -1.50, R) >consensus CGCAGUCAACUAAAUUGCUCCCAGUUAAAAAGCACUUAAGAGGGGUACUUACUCUAGUAUUCCACUUUACGCUAAAAAAAA_GACAUUUCAUCGUCGGCAGGCACGUUUAACAGCUUGUC ....(((.....(((((....)))))....(((....(((.((((((((......)))))))).)))...))).........)))...........(((((((..........))))))) (-25.34 = -25.34 + -0.00)

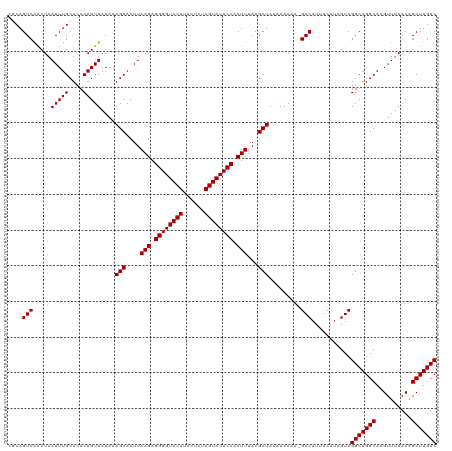
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:24:23 2011