
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 13,577,612 – 13,577,704 |
| Length | 92 |
| Max. P | 0.516443 |

| Location | 13,577,612 – 13,577,704 |
|---|---|
| Length | 92 |
| Sequences | 8 |
| Columns | 106 |
| Reading direction | forward |
| Mean pairwise identity | 79.55 |
| Shannon entropy | 0.39604 |
| G+C content | 0.35224 |
| Mean single sequence MFE | -22.75 |
| Consensus MFE | -9.80 |
| Energy contribution | -10.43 |
| Covariance contribution | 0.63 |
| Combinations/Pair | 1.20 |
| Mean z-score | -2.12 |
| Structure conservation index | 0.43 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.04 |
| SVM RNA-class probability | 0.516443 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
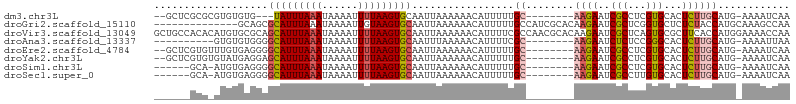
>dm3.chr3L 13577612 92 + 24543557 --GCUCGCGCGUGUGUG---UAUUUAAAUAAAAUUUUAAGUGCAAUUAAAAAACAUUUUUGC--------AAGAAUCGCCUCGUGCACUCUUGCAUG-AAAAUCAA --.((.(((.((((.((---((((((((......))))))))))........))))...)))--------.)).......(((((((....))))))-)....... ( -19.20, z-score = -0.90, R) >droGri2.scaffold_15110 17512086 92 + 24565398 --------------GCAGCGCAUUUAAAUAAAAUUGUAAGUGCAAUUAAAAAACAUUUUUGCCAUCGCACAAGAAUCGCUCGGUGCUCUCUACCAUGCAAAGCCAA --------------((((((((((((.(......).)))))))...........(((((((........))))))).))..((((.....)))).)))........ ( -16.50, z-score = -0.09, R) >droVir3.scaffold_13049 5372658 106 - 25233164 GCUGCCACACAUGUGCGCAGCAUUUAAAUAAAAUUUUAAGUGCAAUUAAAAAACAUUUUCGCCAACGCACAAGAAUCGCUCAGUGCGCUUCACCAUGGAAAACCAA ((((((((....))).)))(((((((((......))))))))).......................))....(((.(((.....))).)))....(((....))). ( -24.80, z-score = -1.74, R) >droAna3.scaffold_13337 1425739 87 - 23293914 ----------GUGUGUGGGGCAUUUAAAUAAAAUUUUAAGUGCAAUUUAAAAACAUUUUCGC--------AAGAAUCUCUCCGGGCACUCUUGCAUG-AAAAUUAA ----------((((.(((((((((((((......)))))))))..............(((..--------..)))....)))).)))).........-........ ( -22.50, z-score = -3.20, R) >droEre2.scaffold_4784 13577687 95 + 25762168 --GCUCGUGUUUGUGAGGGGCAUUUAAAUAAAAUUUUAAGUGCAAUUAAAAAACAUUUUUGC--------AAGAAUCGCCUCGUGCACUCUUGCAUG-AAAAUCAA --((..((((((...((..(((((((((......)))))))))..))...))))))....))--------..........(((((((....))))))-)....... ( -24.20, z-score = -2.20, R) >droYak2.chr3L 13669757 95 + 24197627 --GCUCGUGUGUAUGAGGAGCAUUUAAAUAAAAUUUUAAGUGCAAUUAAAAAACAUUUUUGC--------AAGAAUCGCCUCGUGCACUCUUGCAUG-AAAAUCAA --((..(.((((((((((.(((((((((......)))))))))...(((((.....))))).--------........)))))))))).)..))...-........ ( -27.70, z-score = -3.80, R) >droSim1.chr3L 12956094 90 + 22553184 ------GCA-AUGUGAGGGGCAUUUAAAUAAAAUUUUAAGUGCAAUUAAAAAACAUUUUUGC--------AAGAAUCGCCUCGUGCACUCUUGCAUG-AAAAUCAA ------...-(((..(((((((((((((......)))))))))................(((--------(.((......)).))))))))..))).-........ ( -23.60, z-score = -2.66, R) >droSec1.super_0 5739611 90 + 21120651 ------GCA-AUGUGAGGGGCAUUUAAAUAAAAUUUUAAGUGCAAUUAAAAAACAUUUUUGC--------AAGAAUCGCCUUGUGCACUCUUGCAUG-AAAAUCAA ------...-(((..(((((((((((((......))))))))).................((--------(((......)))))...))))..))).-........ ( -23.50, z-score = -2.38, R) >consensus ______GCA_AUGUGAGGGGCAUUUAAAUAAAAUUUUAAGUGCAAUUAAAAAACAUUUUUGC________AAGAAUCGCCUCGUGCACUCUUGCAUG_AAAAUCAA ...................(((((((((......)))))))))......................................((((((....))))))......... ( -9.80 = -10.43 + 0.63)


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:23:44 2011