| Sequence ID | dm3.chr3L |
|---|---|
| Location | 13,126,320 – 13,126,420 |
| Length | 100 |
| Max. P | 0.689519 |

| Location | 13,126,320 – 13,126,420 |
|---|---|
| Length | 100 |
| Sequences | 8 |
| Columns | 119 |
| Reading direction | forward |
| Mean pairwise identity | 77.66 |
| Shannon entropy | 0.39848 |
| G+C content | 0.56675 |
| Mean single sequence MFE | -33.01 |
| Consensus MFE | -16.74 |
| Energy contribution | -18.18 |
| Covariance contribution | 1.44 |
| Combinations/Pair | 1.20 |
| Mean z-score | -1.94 |
| Structure conservation index | 0.51 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.42 |
| SVM RNA-class probability | 0.689519 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 13126320 100 + 24543557 ------------CACGGGGUCGGGGAGCGGAAGUAGUGGGUU---AAGCCUUGAGGUUAUGGGGAUGACUGG--GCUGGCAAGUCAUC--ACGGACAUUGCACUUUGGCUUUGACCAUC ------------.....(((((..((((.(((((........---.(((((...((((((....))))))))--))).(((((((...--...))).))))))))).)))))))))... ( -34.40, z-score = -2.05, R) >droEre2.scaffold_4784 13134364 112 + 25762168 CAAGGGGUCGGAGUCGGGGUCGGGGAGCGGAAGUAGUGGGUU---AAGCCUUGGGGUUAUGGGGAUGACUGG--GCUGGCAAGUCAUC--ACGGACAUUGCACUUUGGCUUUGACCAUC .....((((((((((((((((.(....).)).((((((....---.((((....)))).....(((((((..--(....).)))))))--.....)))))).))))))))))))))... ( -40.90, z-score = -2.80, R) >droYak2.chr3L 13210916 100 + 24197627 ------------GAGGGGAUCGGGCAGCGGAAGUAGUGGGUU---AAGCCUUGGGGUUAUGGGGAUGACUGG--GCUGGCAAGUCAUC--ACGGACAUUGCACUUUGGCUUUGACCAUC ------------...........((.((....)).)).((((---(((((..((.((.(((..(((((((..--(....).)))))))--.....))).)).))..))).))))))... ( -31.30, z-score = -1.01, R) >droSec1.super_0 5302063 100 + 21120651 ------------CACGGGGUCGGGGAGCGGAAGUAGUGGGUU---AAGCCUUGGGGUUAUGGGGAUGACUGG--GCUGGCAAGUCAUC--ACGGACAUUGCACUUUGGCUUUGACCAUC ------------.....(((((..((((.(((((........---.(((((...((((((....))))))))--))).(((((((...--...))).))))))))).)))))))))... ( -34.40, z-score = -2.02, R) >droSim1.chr3L 12519855 100 + 22553184 ------------CACGGGGUCGGGGAGCGGAAGUAGUGGGUU---AAGCCUUGGGGUUAUGGGGAUGACUGG--GCUGGCAAGUCAUC--ACGGACAUUGCACUUUGGCUUUGACCAUC ------------.....(((((..((((.(((((........---.(((((...((((((....))))))))--))).(((((((...--...))).))))))))).)))))))))... ( -34.40, z-score = -2.02, R) >dp4.chrXR_group6 5394984 95 + 13314419 ------------------CGGGUAUGGGAGGGGGG-UGGCUA---AGGGGGAGGGGCUAUGGGGAUGACUGGCAAGAGGCAAGUCAUC--ACGGACAUUGCACUUUGGCUUUGACCAUC ------------------...............((-(((.((---(((..((((.((.(((..(((((((.((.....)).)))))))--.....))).)).))))..))))).))))) ( -28.70, z-score = -2.04, R) >droPer1.super_484 11380 96 + 20326 ------------------CGGGUAUGGCAGGGGGGGCGGCUU---AGGGGGAGGGGCUAUGGGGAUGACUGGCAAGAGGCAAGUCAUC--ACGGACAUUGCACUUUGGCUUUGACCAUC ------------------..(((..(((..(((..(((((((---........))))).....(((((((.((.....)).)))))))--.........)).)))..)))...)))... ( -29.20, z-score = -1.76, R) >droAna3.scaffold_13337 10945089 96 - 23293914 -------------------UUUUGGGGCGGAAGUAGUGGGUUUGUGGGUUUUCGGAUCU--GGAAUGACUGG--GCUGGCAAGUCAUCUCAUGGACAUUGCACUUUGGCUUUGACCAUC -------------------..(..((((.(((((.(..((((..((((....(((.((.--.....)))))(--(((....))))..))))..))).)..)))))).))))..)..... ( -30.80, z-score = -1.85, R) >consensus _____________ACGGGGUCGGGGAGCGGAAGUAGUGGGUU___AAGCCUUGGGGUUAUGGGGAUGACUGG__GCUGGCAAGUCAUC__ACGGACAUUGCACUUUGGCUUUGACCAUC ..........................((....))....((((...(((((..((.((.(((..(((((((...........))))))).......))).)).))..))))).))))... (-16.74 = -18.18 + 1.44)


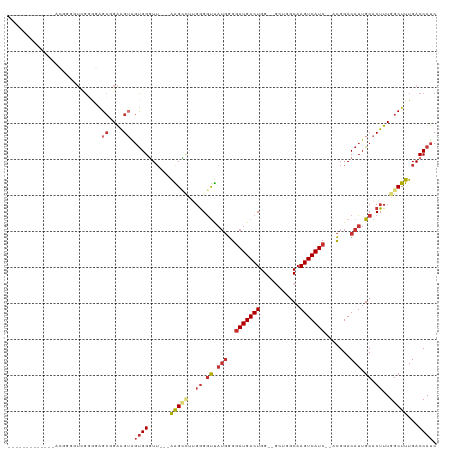
| Location | 13,126,320 – 13,126,420 |
|---|---|
| Length | 100 |
| Sequences | 8 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 77.66 |
| Shannon entropy | 0.39848 |
| G+C content | 0.56675 |
| Mean single sequence MFE | -17.08 |
| Consensus MFE | -12.93 |
| Energy contribution | -12.71 |
| Covariance contribution | -0.22 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.00 |
| Structure conservation index | 0.76 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.31 |
| SVM RNA-class probability | 0.640678 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 13126320 100 - 24543557 GAUGGUCAAAGCCAAAGUGCAAUGUCCGU--GAUGACUUGCCAGC--CCAGUCAUCCCCAUAACCUCAAGGCUU---AACCCACUACUUCCGCUCCCCGACCCCGUG------------ ...((((..(((..(((((..........--(((((((.(.....--).)))))))....(((((....)).))---)......)))))..)))....)))).....------------ ( -18.10, z-score = -0.77, R) >droEre2.scaffold_4784 13134364 112 - 25762168 GAUGGUCAAAGCCAAAGUGCAAUGUCCGU--GAUGACUUGCCAGC--CCAGUCAUCCCCAUAACCCCAAGGCUU---AACCCACUACUUCCGCUCCCCGACCCCGACUCCGACCCCUUG ...((((.(((((.....((.......))--(((((((.(.....--).))))))).............)))))---....................((....)).....))))..... ( -18.90, z-score = -0.89, R) >droYak2.chr3L 13210916 100 - 24197627 GAUGGUCAAAGCCAAAGUGCAAUGUCCGU--GAUGACUUGCCAGC--CCAGUCAUCCCCAUAACCCCAAGGCUU---AACCCACUACUUCCGCUGCCCGAUCCCCUC------------ ...((((..(((..(((((..........--(((((((.(.....--).))))))).............((...---...))..)))))..)))....)))).....------------ ( -16.50, z-score = -0.13, R) >droSec1.super_0 5302063 100 - 21120651 GAUGGUCAAAGCCAAAGUGCAAUGUCCGU--GAUGACUUGCCAGC--CCAGUCAUCCCCAUAACCCCAAGGCUU---AACCCACUACUUCCGCUCCCCGACCCCGUG------------ ...((((..(((..(((((..........--(((((((.(.....--).))))))).............((...---...))..)))))..)))....)))).....------------ ( -18.00, z-score = -0.93, R) >droSim1.chr3L 12519855 100 - 22553184 GAUGGUCAAAGCCAAAGUGCAAUGUCCGU--GAUGACUUGCCAGC--CCAGUCAUCCCCAUAACCCCAAGGCUU---AACCCACUACUUCCGCUCCCCGACCCCGUG------------ ...((((..(((..(((((..........--(((((((.(.....--).))))))).............((...---...))..)))))..)))....)))).....------------ ( -18.00, z-score = -0.93, R) >dp4.chrXR_group6 5394984 95 - 13314419 GAUGGUCAAAGCCAAAGUGCAAUGUCCGU--GAUGACUUGCCUCUUGCCAGUCAUCCCCAUAGCCCCUCCCCCU---UAGCCA-CCCCCCUCCCAUACCCG------------------ .((((.((..((......))..)).))))--(((((((.((.....)).)))))))..................---......-.................------------------ ( -16.90, z-score = -2.35, R) >droPer1.super_484 11380 96 - 20326 GAUGGUCAAAGCCAAAGUGCAAUGUCCGU--GAUGACUUGCCUCUUGCCAGUCAUCCCCAUAGCCCCUCCCCCU---AAGCCGCCCCCCCUGCCAUACCCG------------------ .((((.((..((......))..)).))))--(((((((.((.....)).)))))))..................---........................------------------ ( -16.90, z-score = -1.73, R) >droAna3.scaffold_13337 10945089 96 + 23293914 GAUGGUCAAAGCCAAAGUGCAAUGUCCAUGAGAUGACUUGCCAGC--CCAGUCAUUCC--AGAUCCGAAAACCCACAAACCCACUACUUCCGCCCCAAAA------------------- .((((.((..((......))..)).))))(..((((((.(.....--).))))))..)--........................................------------------- ( -13.30, z-score = -0.24, R) >consensus GAUGGUCAAAGCCAAAGUGCAAUGUCCGU__GAUGACUUGCCAGC__CCAGUCAUCCCCAUAACCCCAAGGCUU___AACCCACUACUUCCGCUCCCCGACCCCGU_____________ .((((.((..((......))..)).))))..(((((((.(.......).)))))))............................................................... (-12.93 = -12.71 + -0.22)



Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:22:50 2011