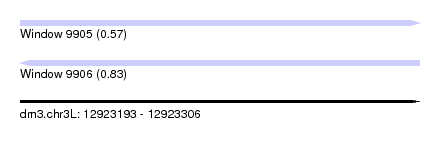

| Sequence ID | dm3.chr3L |
|---|---|
| Location | 12,923,193 – 12,923,306 |
| Length | 113 |
| Max. P | 0.830690 |

| Location | 12,923,193 – 12,923,306 |
|---|---|
| Length | 113 |
| Sequences | 5 |
| Columns | 113 |
| Reading direction | forward |
| Mean pairwise identity | 95.55 |
| Shannon entropy | 0.07689 |
| G+C content | 0.52593 |
| Mean single sequence MFE | -37.53 |
| Consensus MFE | -31.98 |
| Energy contribution | -32.58 |
| Covariance contribution | 0.60 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.76 |
| Structure conservation index | 0.85 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.16 |
| SVM RNA-class probability | 0.569693 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
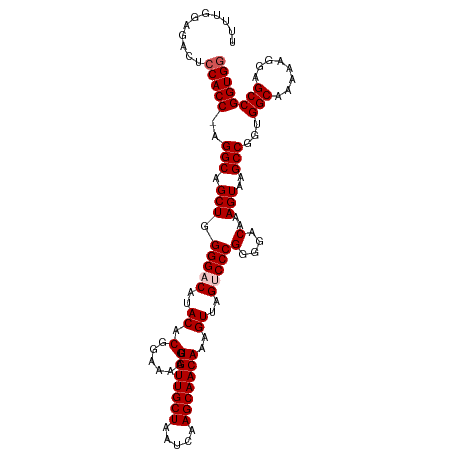
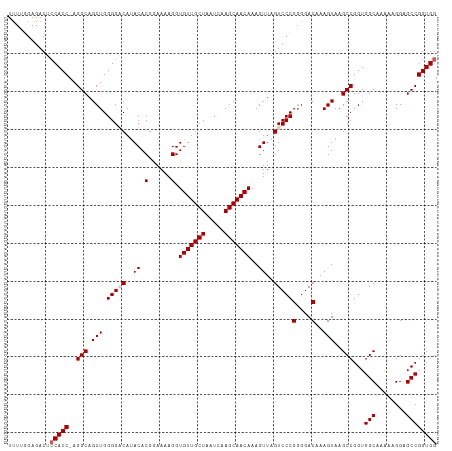
>dm3.chr3L 12923193 113 + 24543557 UUUUGGAGACUCCACCCAGGCAGCUGGGGACAUACACGGAAAAGGUGUUGCUAAUCAAGCAACAAAGUUAGUCCCGUGGACAAAGUAAGCCGGUGGCAAAAAGGAGCCGGUGG ....(....).(((((..(((...((....))..(((((......(((((((.....))))))).........)))))..........)))...(((........)))))))) ( -36.26, z-score = -1.20, R) >droSim1.chr3L 12302457 112 + 22553184 UUUUGGAGACUCCACC-AGGCAGCUGGGGACAUACACGGAAAAGGUGUUGCUAAUCAAGCAACAAAGUUAGUCCCGGGGACAAAGUAAGCCGGUGGCAAAAAGGAGCCGGUGG ....(....).(((((-.(((.(((.(((((..((.(......).(((((((.....)))))))..))..)))))(....)..)))..)))...(((........)))))))) ( -38.40, z-score = -2.13, R) >droSec1.super_0 5098457 112 + 21120651 UCUUGGAGACUCCACC-AGGCAGCUGGGGACAUACACGGAAAAGGUGUUGCUAAUCAAGCAACAAAGUUAGUCCCGGGGACAAAGUAAGCCGGUGGCAAAAAGGAGCCGGUGG ....(....).(((((-.(((.(((.(((((..((.(......).(((((((.....)))))))..))..)))))(....)..)))..)))...(((........)))))))) ( -38.40, z-score = -1.91, R) >droYak2.chr3L 13002913 111 + 24197627 UUUUUGAGACUGCACC-AGGCAGCUGGGG-CAAACACGAGAAAGGUGUUGCUAAUCAAGCAACAAAGUUAGUCCCGGGGACAAAGUAAGCCGGUGGCAAAAUGGAGCCGGUGG .(((((...((((...-..))))((((((-(.(((.(......).(((((((.....)))))))..))).)))))))...)))))...((((((..(.....)..)))))).. ( -37.80, z-score = -1.87, R) >droEre2.scaffold_4784 12932497 111 + 25762168 UUUUUGAGACUCCACC-AGGCAGCUGGGG-CAUACACGAAAAAGGUGUUGCUAAUCAAGCAACAAAGUUAGUCCCGGGGACAAAGUAAGCCGGUGGCAAGAAGGAGCCGGUGG ...........(((((-.(((.(((((((-(..((.(......).(((((((.....)))))))..))..)))))(....)..)))..)))...(((........)))))))) ( -36.80, z-score = -1.67, R) >consensus UUUUGGAGACUCCACC_AGGCAGCUGGGGACAUACACGGAAAAGGUGUUGCUAAUCAAGCAACAAAGUUAGUCCCGGGGACAAAGUAAGCCGGUGGCAAAAAGGAGCCGGUGG ...........(((((..(((.(((.(((((..((.(......).(((((((.....)))))))..))..)))))(....)..)))..)))...(((........)))))))) (-31.98 = -32.58 + 0.60)

| Location | 12,923,193 – 12,923,306 |
|---|---|
| Length | 113 |
| Sequences | 5 |
| Columns | 113 |
| Reading direction | reverse |
| Mean pairwise identity | 95.55 |
| Shannon entropy | 0.07689 |
| G+C content | 0.52593 |
| Mean single sequence MFE | -32.22 |
| Consensus MFE | -29.62 |
| Energy contribution | -29.82 |
| Covariance contribution | 0.20 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.72 |
| Structure conservation index | 0.92 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.83 |
| SVM RNA-class probability | 0.830690 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
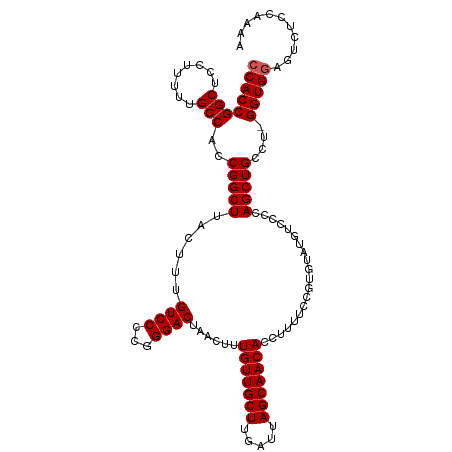
>dm3.chr3L 12923193 113 - 24543557 CCACCGGCUCCUUUUUGCCACCGGCUUACUUUGUCCACGGGACUAACUUUGUUGCUUGAUUAGCAACACCUUUUCCGUGUAUGUCCCCAGCUGCCUGGGUGGAGUCUCCAAAA ((((((((........)))..(((((.....(((.(((((((.......(((((((.....)))))))....))))))).))).....)))))....)))))........... ( -36.00, z-score = -2.44, R) >droSim1.chr3L 12302457 112 - 22553184 CCACCGGCUCCUUUUUGCCACCGGCUUACUUUGUCCCCGGGACUAACUUUGUUGCUUGAUUAGCAACACCUUUUCCGUGUAUGUCCCCAGCUGCCU-GGUGGAGUCUCCAAAA ((((((((..((....(((...))).............(((((.......((((((.....))))))(((......).))..))))).))..))).-)))))........... ( -31.30, z-score = -1.61, R) >droSec1.super_0 5098457 112 - 21120651 CCACCGGCUCCUUUUUGCCACCGGCUUACUUUGUCCCCGGGACUAACUUUGUUGCUUGAUUAGCAACACCUUUUCCGUGUAUGUCCCCAGCUGCCU-GGUGGAGUCUCCAAGA ((((((((..((....(((...))).............(((((.......((((((.....))))))(((......).))..))))).))..))).-)))))........... ( -31.30, z-score = -1.26, R) >droYak2.chr3L 13002913 111 - 24197627 CCACCGGCUCCAUUUUGCCACCGGCUUACUUUGUCCCCGGGACUAACUUUGUUGCUUGAUUAGCAACACCUUUCUCGUGUUUG-CCCCAGCUGCCU-GGUGCAGUCUCAAAAA .....(((........)))...(((.......)))...((((((.....(((((((.....)))))))..............(-(.((((....))-)).))))))))..... ( -29.90, z-score = -1.38, R) >droEre2.scaffold_4784 12932497 111 - 25762168 CCACCGGCUCCUUCUUGCCACCGGCUUACUUUGUCCCCGGGACUAACUUUGUUGCUUGAUUAGCAACACCUUUUUCGUGUAUG-CCCCAGCUGCCU-GGUGGAGUCUCAAAAA ((((((((........)))...(((.(((...((((...))))......(((((((.....)))))))..........))).)-))..........-)))))........... ( -32.60, z-score = -1.91, R) >consensus CCACCGGCUCCUUUUUGCCACCGGCUUACUUUGUCCCCGGGACUAACUUUGUUGCUUGAUUAGCAACACCUUUUCCGUGUAUGUCCCCAGCUGCCU_GGUGGAGUCUCCAAAA ((((((((........)))..(((((......((((...))))......(((((((.....)))))))....................)))))....)))))........... (-29.62 = -29.82 + 0.20)


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:22:18 2011