
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 12,824,707 – 12,824,802 |
| Length | 95 |
| Max. P | 0.842191 |

| Location | 12,824,707 – 12,824,802 |
|---|---|
| Length | 95 |
| Sequences | 7 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 67.97 |
| Shannon entropy | 0.57236 |
| G+C content | 0.38753 |
| Mean single sequence MFE | -22.14 |
| Consensus MFE | -10.45 |
| Energy contribution | -9.74 |
| Covariance contribution | -0.71 |
| Combinations/Pair | 1.58 |
| Mean z-score | -1.47 |
| Structure conservation index | 0.47 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.88 |
| SVM RNA-class probability | 0.842191 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 12824707 95 - 24543557 -----UUUUUUUCUCAGUGCUUGCU------------UAACUUUAGUUAAUUAACGCUGUCCAGCUG-----GUUAGCCAAGUUAUAAUGCAUCUUUUGAUG-UUGACCUUGGCUGAG -----.........(((((.(((.(------------((((....))))).)))))))).((....)-----)((((((((((((....(((((....))))-)))).))))))))). ( -26.30, z-score = -2.78, R) >droAna3.scaffold_13337 3532771 115 + 23293914 CAAAAUAUUUUUCUCUAUGUAUGCUUAUGGUGCCGGCUAAUUUAAGUUAAUUUGUGCUUCCCAUCUGGCCUAGCUAGCCAAGU--UAAUACAUCCUCCGGUA-UUGACCUUGACUAUA ......................(((..(((.(((((.......((((........)))).....))))))))...)))(((((--((((((........)))-)))).))))...... ( -22.00, z-score = -0.36, R) >droEre2.scaffold_4784 12836889 95 - 25762168 -----UAUUUUUCUCAGUGCUUGCU------------UAACUUAAGUAAAUUAACGUUGUCCAGCUG-----GCUAGCCAGGUCUUAAUGCAUUUUUUGAUG-UUGACCUUGGCUAAG -----.............(((.(((------------((((((((.....)))).))))...))).)-----))((((((((((.....(((((....))))-).)))).)))))).. ( -25.60, z-score = -2.27, R) >droYak2.chr3L 12899877 93 - 24197627 -------UUUUUCUCAGUGCUUGCU------------UAACUUAAGUAAUUUAACGCUGACCAGCUG-----GCUAGCCAAGUUAUAAUGCAUCUUUUGAUG-UUGACCUUGGUUGAG -------......((((((.(((((------------((...))))))).....))))))((....)-----).(((((((((((....(((((....))))-)))).)))))))).. ( -22.30, z-score = -1.26, R) >droSec1.super_0 5008797 97 - 21120651 ---UAUUUUUUUCUCAGUGCUUGCU------------UAACUUUAGUUAAUUAACGCUGUCCAGCUG-----GCUAGCCAAGUUAUAAUGCAUCUUUUGAUG-UUGACCUUGGUUGAG ---...........(((((.(((.(------------((((....))))).))))))))........-----..(((((((((((....(((((....))))-)))).)))))))).. ( -22.30, z-score = -1.31, R) >droSim1.chr3L 12211083 96 - 22553184 ----UAUUUUUUCUCAGUGCUUGCU------------UAACUUUAGUUAAUAAACGCUGUCCAGCUG-----GCUAGCCAAGUUAUAAUGCAUCUUUUGAUG-UUGACCUUGGCUGAG ----........(((...(((.(((------------..((....(((....)))...))..))).)-----)).((((((((((....(((((....))))-)))).)))))))))) ( -24.70, z-score = -1.82, R) >droWil1.scaffold_180698 5939087 84 + 11422946 -------CCUUUCCC----------------------CCACUCACCCUAAAAAAGAAGUCGAAACCA-----AUUAGCGCAGUGCGAAAUCAUUUUUGUAUGCUCGAUAUGCGUUGUU -------........----------------------..............................-----..(((((((...(((.((((....)).))..)))...))))))).. ( -11.80, z-score = -0.52, R) >consensus _____UUUUUUUCUCAGUGCUUGCU____________UAACUUAAGUUAAUUAACGCUGUCCAGCUG_____GCUAGCCAAGUUAUAAUGCAUCUUUUGAUG_UUGACCUUGGCUGAG ..........................................................................((((((((........((((....))))......)))))))).. (-10.45 = -9.74 + -0.71)


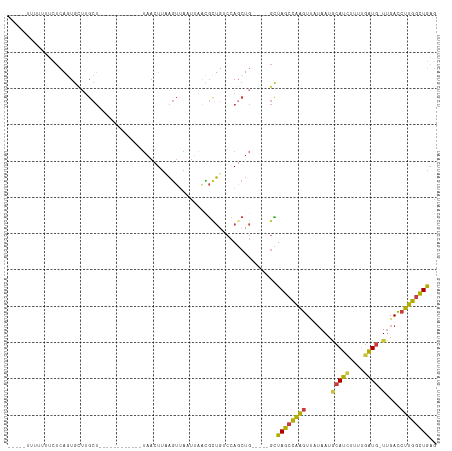
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:22:09 2011