
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 12,618,796 – 12,618,890 |
| Length | 94 |
| Max. P | 0.669609 |

| Location | 12,618,796 – 12,618,890 |
|---|---|
| Length | 94 |
| Sequences | 3 |
| Columns | 116 |
| Reading direction | reverse |
| Mean pairwise identity | 75.70 |
| Shannon entropy | 0.31232 |
| G+C content | 0.49588 |
| Mean single sequence MFE | -33.47 |
| Consensus MFE | -20.13 |
| Energy contribution | -20.13 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.00 |
| Structure conservation index | 0.60 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.38 |
| SVM RNA-class probability | 0.669609 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 12618796 94 - 24543557 --------------GACUUCGAGUUUU--AAAUGCGAUCUGUGGCGCAGGUAUCCGCGAGAUACUU-----GCGCUGC-UGCCACAUAAUGGUAUGGUAUAUUCUGCCGCCAACUG --------------.............--....((.....(..(((((((((((.....)))))))-----))))..)-(((((.....))))).((((.....))))))...... ( -30.00, z-score = -1.53, R) >droYak2.chr3L 12684393 111 - 24197627 GAGUUCGAGUUUAGCAAUGCGAAAUGCGAAAAUGCGAUCUGUGGCGCAGGUAUCCGCAAGAUACUUGCCUCGCGCUGCUUGCCACAUAAUGGU--GGUAUAUGCU---GCCAACUG .((.(((.((((.(((........)))..)))).))).))((((.((((((((.(....))))))))).))))((.((.((((((......))--))))...)).---))...... ( -36.60, z-score = -1.00, R) >droEre2.scaffold_4784 12623703 89 - 25762168 --------------GAGUUCGAGCUUU--AAAUGUGAUCUGUGGCGCAGGUAUCCACGAGAUACUU-----GCGCUGC-UGCCACAUAAUGGU--GGUAUAUUCU---GCCAACUG --------------((((....)))).--..(((((....(..(((((((((((.....)))))))-----))))..)-...)))))..(((.--.(......).---.))).... ( -33.80, z-score = -3.46, R) >consensus ______________GAAUUCGAGAUUU__AAAUGCGAUCUGUGGCGCAGGUAUCCGCGAGAUACUU_____GCGCUGC_UGCCACAUAAUGGU__GGUAUAUUCU___GCCAACUG ...............................(((.(....(..(((((((((((.....))))))).....))))..)....).)))..((((..((......))...)))).... (-20.13 = -20.13 + 0.00)

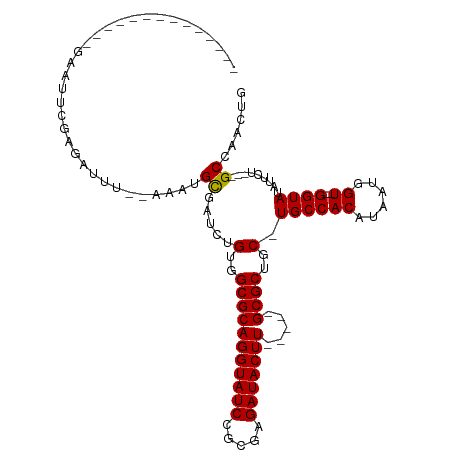
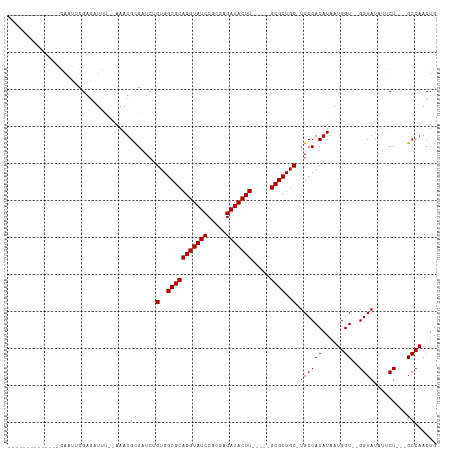
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:21:38 2011