
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 12,570,486 – 12,570,545 |
| Length | 59 |
| Max. P | 0.557184 |

| Location | 12,570,486 – 12,570,545 |
|---|---|
| Length | 59 |
| Sequences | 12 |
| Columns | 59 |
| Reading direction | forward |
| Mean pairwise identity | 96.45 |
| Shannon entropy | 0.07752 |
| G+C content | 0.34221 |
| Mean single sequence MFE | -8.28 |
| Consensus MFE | -8.28 |
| Energy contribution | -8.28 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -0.73 |
| Structure conservation index | 1.00 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.13 |
| SVM RNA-class probability | 0.557184 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
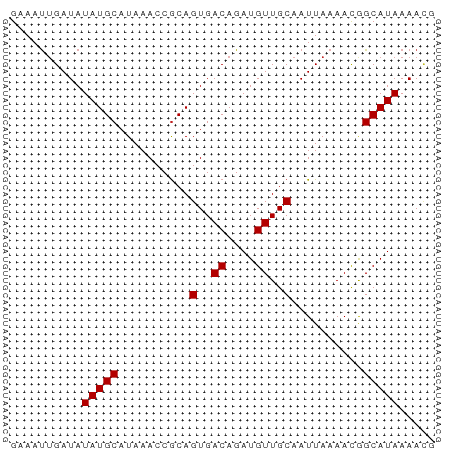
>dm3.chr3L 12570486 59 + 24543557 GAAAUUGAUAUAUGCAUAAACCGCAGUGACAGAUGUUGCAAUUAAAACGGCAUAAAACG ..........(((((..........(..((....))..)..........)))))..... ( -8.65, z-score = -0.70, R) >droSim1.chr3L 11954632 59 + 22553184 GAAAUUGAUAUAUGCAUAAACCGCAGUGACAGAUGUUGCAAUUAAAACGGCAUAAAACG ..........(((((..........(..((....))..)..........)))))..... ( -8.65, z-score = -0.70, R) >droSec1.super_0 4756905 59 + 21120651 GAAAUUGAUAUAUGCAUAAACCGCAGUGACAGAUGUUGCAAUUAAAACGGCAUAAAACG ..........(((((..........(..((....))..)..........)))))..... ( -8.65, z-score = -0.70, R) >droYak2.chr3L 12633858 59 + 24197627 GAAAUUGAUAUAUGCAUAAACCGCAGUGACAGAUGUUGCAAUUAAAACGGCAUAAAACG ..........(((((..........(..((....))..)..........)))))..... ( -8.65, z-score = -0.70, R) >droEre2.scaffold_4784 12572971 59 + 25762168 GAAAUUGAUAUAUGCAUAAACCGCAGUGACAGAUGUUGCAAUUAAAACGGCAUAAAACG ..........(((((..........(..((....))..)..........)))))..... ( -8.65, z-score = -0.70, R) >droAna3.scaffold_13337 19839473 59 - 23293914 GAAAUUGAUAUAUGCAUAAACCGCAGUGACAGAUGUUGCAAUUAAAACGGCAUAAAACG ..........(((((..........(..((....))..)..........)))))..... ( -8.65, z-score = -0.70, R) >dp4.chrXR_group6 3354928 59 + 13314419 GAAAUUGAUAUAUGCAUAAACCGCAGUGACAGAUGUUGCAAUUAAAACAGCAUAAAACG ..........(((((..........(..((....))..)..........)))))..... ( -7.55, z-score = -0.56, R) >droPer1.super_9 1653253 59 + 3637205 GAAAUUGAUAUAUGCAUAAACCGCAGUGACAGAUGUUGCAAUUAAAACAGCAUAAAACG ..........(((((..........(..((....))..)..........)))))..... ( -7.55, z-score = -0.56, R) >droWil1.scaffold_181136 2221616 59 - 2313701 GAAACCGAUAUAUGCAUAAACCACAGUGACAGAUGUUGCAAUUAAAAUAGCAUAAAAUG ..........(((((..........(..((....))..)..........)))))..... ( -7.55, z-score = -1.41, R) >droMoj3.scaffold_6680 17763944 57 - 24764193 GAAAUUGAUAUAUGCAUAAACCGCAGUGACAGAUGUUGCAAUUAAAACAGCAUAAAA-- ..........(((((..........(..((....))..)..........)))))...-- ( -7.55, z-score = -0.48, R) >droGri2.scaffold_15110 7618280 58 - 24565398 GAAAUUGAUAUAUGCAUAAACCGCAGUGACAGAUGUUGCAAUUAAAACGGCAUAAAAU- ..........(((((..........(..((....))..)..........)))))....- ( -8.65, z-score = -0.76, R) >droVir3.scaffold_13049 5969789 58 - 25233164 GAAAUUGAUAUAUGCAUAAACCGCAGUGACAGAUGUUGCAAUUAAAACGGCAUAAAAU- ..........(((((..........(..((....))..)..........)))))....- ( -8.65, z-score = -0.76, R) >consensus GAAAUUGAUAUAUGCAUAAACCGCAGUGACAGAUGUUGCAAUUAAAACGGCAUAAAACG ..........(((((..........(..((....))..)..........)))))..... ( -8.28 = -8.28 + -0.00)


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:21:26 2011