
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 12,413,146 – 12,413,240 |
| Length | 94 |
| Max. P | 0.749591 |

| Location | 12,413,146 – 12,413,240 |
|---|---|
| Length | 94 |
| Sequences | 10 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 71.55 |
| Shannon entropy | 0.53641 |
| G+C content | 0.42878 |
| Mean single sequence MFE | -17.52 |
| Consensus MFE | -7.70 |
| Energy contribution | -7.96 |
| Covariance contribution | 0.26 |
| Combinations/Pair | 1.29 |
| Mean z-score | -1.64 |
| Structure conservation index | 0.44 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.58 |
| SVM RNA-class probability | 0.749591 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 12413146 94 - 24543557 ---------------GACCAGGAGAAUAAGUAAUUCACUUAACGCUUUAAUAUCGCUGAUAAAU--GCAGACCAAUCUGCAUUCCCC---CCCUUUGAUUUCUGUUCUUUUUUU ---------------((.((((((..(((((.....)))))....................(((--(((((....))))))))....---........)))))).))....... ( -16.40, z-score = -2.02, R) >droSim1.chr3L 11793952 97 - 22553184 ---------------GACCAGGAGAAUAAGUAAUUCACUUAACGCUUUAAUAUCGCUGAUAAAU--GCAGACCAAUCUGCAUUCCCCCCACCAUUCGAUUUCUGUUCUUUUUUU ---------------((.((((((..(((((.....)))))....................(((--(((((....))))))))...............)))))).))....... ( -16.40, z-score = -2.08, R) >droSec1.super_0 4597032 97 - 21120651 ---------------GACCAGGAGAAUAAGUAAUUCACUUAACGCUUUAAUAUCGCUGAUAAAU--GCAGACCAAUCUGCAUUCCCCCCUCCCUUCGAUUUCUGUUCUUUUUUU ---------------((.((((((..(((((.....)))))....................(((--(((((....))))))))...............)))))).))....... ( -16.40, z-score = -2.02, R) >droYak2.chr3L 12469368 93 - 24197627 ---------------GACCAGGAGAAUAAGUAAUUCACUUAACGCUUUAAUAUCGCUGAUAAAU--GCAGACCAAUCUGCAUUCCC----CCCUCUGAUUUCAGUUCCCUUUUU ---------------....(((.((.(((((.....))))).............(((((..(((--(((((....))))))))...----..........)))))))))).... ( -15.86, z-score = -1.80, R) >droEre2.scaffold_4784 12408321 87 - 25762168 ---------------GACCAGGAGAAUAAGUAAUUCACUUAACGCUUUAAUAUCGCUGAUAAAU--GCAGACCAAUCUGCAUUCCC----CCCUCCGAUCUCGGUUCC------ ---------------((((.((((..(((((.....)))))....................(((--(((((....))))))))...----..))))(....)))))..------ ( -19.10, z-score = -2.53, R) >dp4.chrXR_group6 3198042 82 - 13314419 ---------------GGCGACGGGAAUAAGUAAUUCACUUAACGCUUUAAUAUCGCUGAUAAAU--GCAGAGCAAUCUGC---UC-----CCUCCAG-CUCCAGCUCC------ ---------------(((....(((.(((((.....))))).............((((......--(((((....)))))---..-----....)))-)))).)))..------ ( -21.40, z-score = -1.87, R) >droPer1.super_9 1492623 82 - 3637205 ---------------GGCGACGGGAAUAAGUAAUUCACUUAACGCUUUAAUAUCGCUGAUAAAU--GCAGAGCAAUCUGC---UC-----CCUCCAG-CUCCAGCUCC------ ---------------(((....(((.(((((.....))))).............((((......--(((((....)))))---..-----....)))-)))).)))..------ ( -21.40, z-score = -1.87, R) >droWil1.scaffold_180916 991049 101 + 2700594 UGCUUUUGCUGUAGCGGCAAGGAGAAUAAGUAAUUCACUUAACGCUUUAAUAUCUCUGAUAAAU--GCAGGCCAAUCUGCAGGGC-----CCUCUAGACCUUAGCAGC------ ((((....(((.((.(((..(((((..((((............)))).....)))))......(--(((((....))))))..))-----))).))).....))))..------ ( -23.40, z-score = 0.81, R) >droVir3.scaffold_13049 15896771 83 + 25233164 -------------GGCCUAUAGAGAAUAAGUAAUUCACUUAACGCUUUAAUAUCGCUGAUAAAU--GCGAAGCAAU-------GC-----CCCCAGGAUCCGCACUCCCC---- -------------(((...(((((..(((((.....)))))...)))))...((((........--))))......-------))-----)....(((.......)))..---- ( -15.20, z-score = -0.92, R) >droGri2.scaffold_15110 19454308 77 + 24565398 -----------------CACAGAGAAUAAGUAAUUCACUUAACGCUUUAAUAUCGCUGAUAAAUACGCAGCCCAAGCAUCUUCCCCCCACUCCC-------------------- -----------------....(((..(((((.....)))))..((((.......((((.........))))..))))............)))..-------------------- ( -9.60, z-score = -2.13, R) >consensus _______________GACCAGGAGAAUAAGUAAUUCACUUAACGCUUUAAUAUCGCUGAUAAAU__GCAGACCAAUCUGCAUUCCC____CCCCCAGAUUUCAGUUCC______ ....................((((..(((((.....)))))...))))..................(((((....))))).................................. ( -7.70 = -7.96 + 0.26)

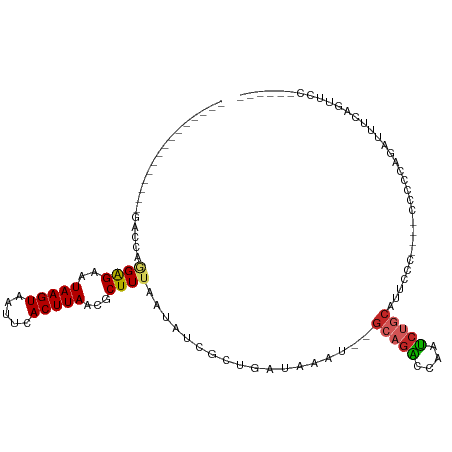
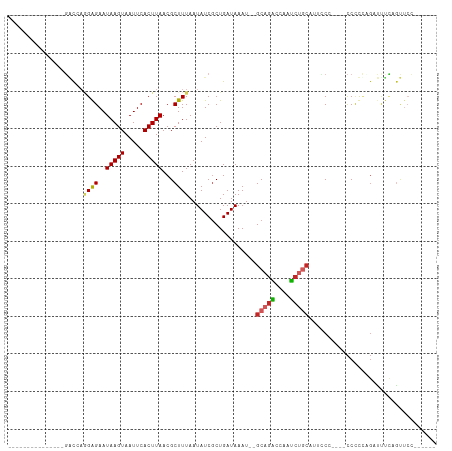
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:21:04 2011