| Sequence ID | dm3.chr3L |
|---|---|
| Location | 12,373,639 – 12,373,709 |
| Length | 70 |
| Max. P | 0.966773 |

| Location | 12,373,639 – 12,373,709 |
|---|---|
| Length | 70 |
| Sequences | 5 |
| Columns | 70 |
| Reading direction | reverse |
| Mean pairwise identity | 84.29 |
| Shannon entropy | 0.25327 |
| G+C content | 0.49186 |
| Mean single sequence MFE | -19.26 |
| Consensus MFE | -15.81 |
| Energy contribution | -16.29 |
| Covariance contribution | 0.48 |
| Combinations/Pair | 1.12 |
| Mean z-score | -2.24 |
| Structure conservation index | 0.82 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.77 |
| SVM RNA-class probability | 0.966773 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
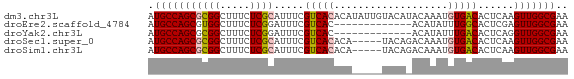
>dm3.chr3L 12373639 70 - 24543557 AUGCCAGCGCGGCUUUCUCGCAUUUCGUCACACAUAUUGUACAUACAAAUGUGACACUCAAGUUGGCGAA .(((((((((((.....)))).....((((((....((((....)))).))))))......))))))).. ( -23.50, z-score = -3.47, R) >droEre2.scaffold_4784 12368953 57 - 25762168 AUGCCAGCGUGGCUUUCUCGGAUUUCGUCAC-------------ACAUAUUUGGCACUCGAGUUGGCGAA .((((((((((((.............)))))-------------.((....))........))))))).. ( -15.52, z-score = -0.93, R) >droYak2.chr3L 12429012 57 - 24197627 AUGCCAGCGCGGCUUUCUCGGAUUUCGUCAC-------------ACAUAUUUGACACUCAGGUUGGCGAA .(((((((.(((.....)))......((((.-------------.......))))......))))))).. ( -14.30, z-score = -0.97, R) >droSec1.super_0 4557748 65 - 21120651 AUGCCAGCGCGGCUUUCUCGCAUUUCGUCACACA-----UACAGACAAAUGUGACACUCAAGUUGGCGAA .(((((((((((.....)))).....((((((..-----..........))))))......))))))).. ( -21.50, z-score = -2.91, R) >droSim1.chr3L 11754121 65 - 22553184 AUGCCAGCGCGGCUUUCUCGCAUUUCGUCACACA-----UACAGACAAAUGUGACACUCAAGUUGGCGAA .(((((((((((.....)))).....((((((..-----..........))))))......))))))).. ( -21.50, z-score = -2.91, R) >consensus AUGCCAGCGCGGCUUUCUCGCAUUUCGUCACACA_____UACA_ACAAAUGUGACACUCAAGUUGGCGAA .(((((((((((.....)))).....(((((...................)))))......))))))).. (-15.81 = -16.29 + 0.48)
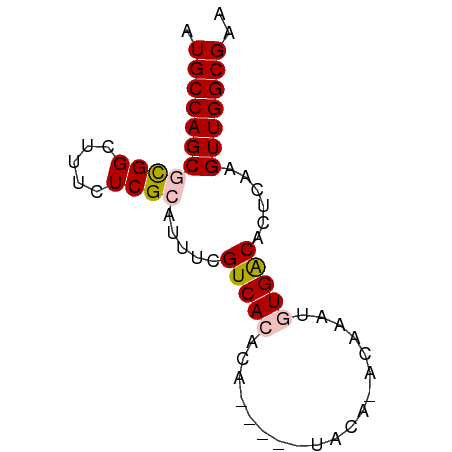
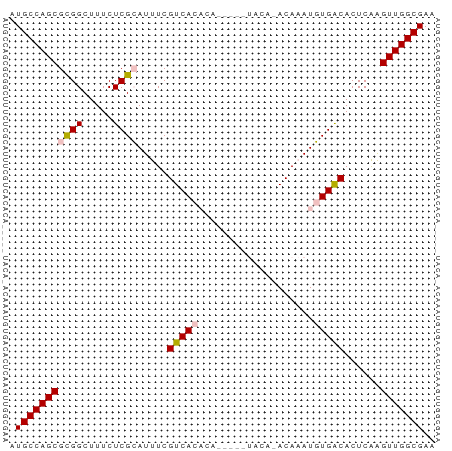
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:20:53 2011