
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 12,295,866 – 12,295,965 |
| Length | 99 |
| Max. P | 0.522916 |

| Location | 12,295,866 – 12,295,965 |
|---|---|
| Length | 99 |
| Sequences | 10 |
| Columns | 108 |
| Reading direction | reverse |
| Mean pairwise identity | 78.70 |
| Shannon entropy | 0.38009 |
| G+C content | 0.49711 |
| Mean single sequence MFE | -26.70 |
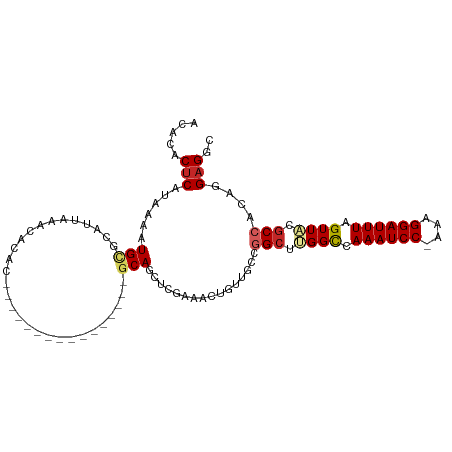
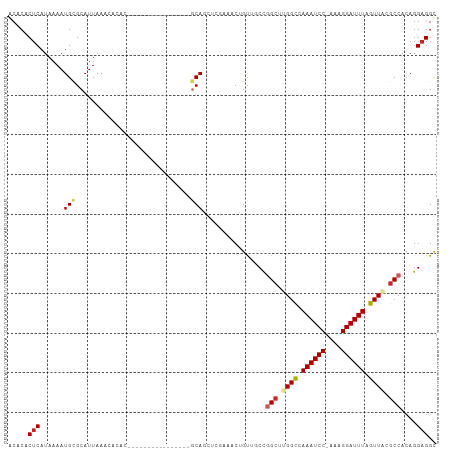
| Consensus MFE | -13.04 |
| Energy contribution | -13.46 |
| Covariance contribution | 0.42 |
| Combinations/Pair | 1.16 |
| Mean z-score | -1.94 |
| Structure conservation index | 0.49 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.06 |
| SVM RNA-class probability | 0.522916 |
| Prediction | RNA |
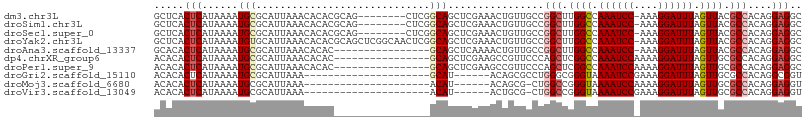
Download alignment: ClustalW | MAF
>dm3.chr3L 12295866 99 - 24543557 GCUCACUCAUAAAAUGCGCAUUAAACACACGCAG--------CUCGGCAGCUCGAAACUGUUGCCGGCUUGGCCAAAUCC-AAAGGAUUUAGUUACGCCACAGGAGGC ..............((((...........)))).--------(((((((((........))))))(((.((((.((((((-...)))))).)))).)))....))).. ( -29.80, z-score = -1.72, R) >droSim1.chr3L 11674078 99 - 22553184 GCUCACUCAUAAAAUGCGCAUUAAACACACGCAG--------CUCGGCAGCUCGAAACUGUUGCCGGCUUGGCCAAAUCC-AAAGGAUUUAGUUACGCCACAGGAGGC ..............((((...........)))).--------(((((((((........))))))(((.((((.((((((-...)))))).)))).)))....))).. ( -29.80, z-score = -1.72, R) >droSec1.super_0 4480530 99 - 21120651 GCUCACUCAUAAAAUGCGCAUUAAACACACGCAG--------CUCGGCAGCUCGAAACUGUUGCCGGCUUGGCCAAAUCC-AAAGGAUUUAGUUACGCCACAGGAGGC ..............((((...........)))).--------(((((((((........))))))(((.((((.((((((-...)))))).)))).)))....))).. ( -29.80, z-score = -1.72, R) >droYak2.chr3L 12348324 107 - 24197627 GCUCACUCAUAAAAUGUGCAUUAAACACACGCAGCUCGGCAACUCGGCAGCUCGAAACUGUUGCCGGCUUGGCCAAAUCC-AAAGGAUUUAGUUACGCCACAGGAGGC ..............((((.......)))).((......))..(((((((((........))))))(((.((((.((((((-...)))))).)))).)))....))).. ( -30.40, z-score = -1.06, R) >droAna3.scaffold_13337 21941564 91 + 23293914 GCACACUCAUAAAAUGCGCAUUAAACACAC----------------GCAGCUCAAAACUGUUGCCGGCUUGGCCAAAUCC-AAAGGAUUUAGUUACGCCACAGGAGGC .....(((......................----------------(((((........))))).(((.((((.((((((-...)))))).)))).)))....))).. ( -22.90, z-score = -1.20, R) >dp4.chrXR_group6 2647884 92 - 13314419 ACACACUCAUAAAAUGCGCAUUAAACACAC----------------GCAGCUCGAAGCCGUUCCCAGCUCGGCCAAAUCCAAAAGGAUUUAGUUGCGCCACAGGAGGC .....(((.......(((((.........(----------------(.((((.(((....)))..))))))((.((((((....)))))).))))))).....))).. ( -21.60, z-score = -0.96, R) >droPer1.super_9 942598 92 - 3637205 ACACACUCAUAAAAUGCGCAUUAAACACAC----------------GCAGCUCGAAGCCGUUCCCAGCUCGGCCAAAUCCAAAAGGAUUUAGUUGCGCCACAGGAGGC .....(((.......(((((.........(----------------(.((((.(((....)))..))))))((.((((((....)))))).))))))).....))).. ( -21.60, z-score = -0.96, R) >droGri2.scaffold_15110 19342082 81 + 24565398 ACACACUCAUAAAAUGCGCAUUAAA---------------------GCAU------ACAGCGCCUGGGCGGGUAAAAUCCGAAAGGAUUUAGUUGCGCCACAGGCGGU .............((((........---------------------))))------....((((((((((.(..((((((....))))))...).)))).)))))).. ( -33.30, z-score = -4.81, R) >droMoj3.scaffold_6680 5906425 80 - 24764193 ACACACUCAUAAAAUGCGCAUUAAA---------------------ACAU------ACAGCG-CUGGCCGGGUAAAAUCCAAAAGGAUUUAGUUGCGCCACAGGAGGU .....(((.......((((......---------------------....------...)))-)(((((((.((((.(((....))))))).))).))))...))).. ( -21.22, z-score = -1.95, R) >droVir3.scaffold_13049 7652667 80 - 25233164 ACACACUCAUAAAAUGCGCAUUAAA---------------------ACAU------ACUGCG-CUGGCCGGGUAAAAUCCGAAAGGAUUUAGUUGCGCCACAGGAGGU .....(((.......(((((.((..---------------------...)------).))))-)(((((((.((((.(((....))))))).))).))))...))).. ( -26.60, z-score = -3.28, R) >consensus ACACACUCAUAAAAUGCGCAUUAAACACAC________________GCAGCUCGAAACUGUUGCCGGCUUGGCCAAAUCC_AAAGGAUUUAGUUACGCCACAGGAGGC .....(((......(((.............................)))................(((.((((.((((((....)))))).)))).)))....))).. (-13.04 = -13.46 + 0.42)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:20:46 2011