
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 12,280,461 – 12,280,573 |
| Length | 112 |
| Max. P | 0.710628 |

| Location | 12,280,461 – 12,280,573 |
|---|---|
| Length | 112 |
| Sequences | 9 |
| Columns | 119 |
| Reading direction | forward |
| Mean pairwise identity | 76.94 |
| Shannon entropy | 0.46629 |
| G+C content | 0.45758 |
| Mean single sequence MFE | -25.08 |
| Consensus MFE | -15.94 |
| Energy contribution | -15.16 |
| Covariance contribution | -0.79 |
| Combinations/Pair | 1.27 |
| Mean z-score | -1.13 |
| Structure conservation index | 0.64 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.48 |
| SVM RNA-class probability | 0.710628 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 12280461 112 + 24543557 ----UGUCUCCAUUUGGCAUUCAAAAACUGUGCGUCUGUGCGAUUUUGACGCAUAUUUGCCGGGCCCAAAACUACCUCACCCAAACUGUC---GUUCGCAUUGAUACUCACAUUGGUAC ----.....(((..((((((.........))))(((.((((((...(((((((....))).(((...............))).....)))---).)))))).))).....)).)))... ( -25.86, z-score = -0.85, R) >droEre2.scaffold_4784 12275667 112 + 25762168 ----UGUCUCCAUUUGGCAUUCAAAAACUGUGCGUCUGUGCGAUUUUGACGCAUAUUUGCCGGGCCCAAAACUACCUCACCCAAACUCUC---AUUCUCAUUCAUACUCGCAUUGGUAC ----.....(((...((((.........((((((((...........))))))))..))))(((...............)))........---....................)))... ( -19.06, z-score = 0.03, R) >droYak2.chr3L 12332385 112 + 24197627 ----UGUCUCCAUUUGGCAUUCAAAAACUGUGCGUCUGUGCGAUUUUGACGCAUAUAUGCCGGGCCCAAAACUACCUCUCCCAAACUCCC---AUUCGCAUUCAUACUCACAUUGGUAC ----.....(((...(((((........((((((((...........)))))))).)))))(((...............)))........---....................)))... ( -19.76, z-score = -0.05, R) >droSec1.super_0 4465196 112 + 21120651 ----UGUCUCCAUUUGGCAUUCAAAAACUGUGCGUCUGUGCGAUUUUGACGCAUAUUUGCCGGGCCCAAAACUACCUCACCCAAACUGUC---GUUCGCAUUGAUACUCACAUGGGUAC ----.((..((((.((((((.........))))(((.((((((...(((((((....))).(((...............))).....)))---).)))))).)))...)).))))..)) ( -28.96, z-score = -1.35, R) >droSim1.chr3L 11658831 112 + 22553184 ----UGUCUCCAUUUGGCAUUCAAAAACUGUGCGUCUGUGCGAUUUUGACGCAUAUUUGCCGGGCCCAAAACUACCUCACCCAAACUGUC---GUUCGCAUUGAUACUCACAUGGUUAC ----.((..((((.((((((.........))))(((.((((((...(((((((....))).(((...............))).....)))---).)))))).)))...)).))))..)) ( -28.26, z-score = -1.63, R) >droAna3.scaffold_13337 21928850 119 - 23293914 UGUGUGUCUCCAUUUGGCAUUCAAAAACUGUGCGUCUGUGCGAUUUUGACGCAUAUUUGCCAGGCCCAGAACCCUCUGGCCCACAGCCUCUUAACAAUCAUUGAUGCGAACAUCUGGCC ((.(((((.......))))).)).....((((((((...........))))))))...((((((.(((((....))))).))....................((((....)))))))). ( -32.70, z-score = -1.14, R) >dp4.chrXR_group6 2629876 107 + 13314419 ----UGUCUCCAUUUGGUAUUCAAAAACUGUGCGUCUGUGCGAUUUUGACGCAUAUUUGCCAAGCACACAGCCAGCACACACAGU--------AUGACCAUCUAUACUCGAAUCCCCAU ----...........((.((((......(((((..(((((.(...(((..(((....)))))).).)))))...)))))...(((--------(((......)))))).)))).))... ( -24.00, z-score = -1.34, R) >droPer1.super_9 925055 97 + 3637205 ----UGUCUCCAUUUGGUAUUCAAAAACUGUGCGUCUGUGCGAUUUUGACGCAUAUUUGCCAAGCACACAGCC----------GU--------AUGACCAUCUAUACUCGAAUCCCCAU ----...........((.((((.....(((((.(..((.((((...((...))...))))))..).)))))..----------((--------(((......)))))..)))).))... ( -20.10, z-score = -1.02, R) >droGri2.scaffold_15110 19322956 88 - 24565398 ----AGUCUCCAUUUGGUAUUCAAAAACUGUGCGUCUGUGUGAUUU-GACGCAUAUUUGCCAAACAUACA------------------------CAGGCAUACAUAC--AAAUGGGCAC ----.((.((((((((((((.........))))(((((((((.(((-(..(((....)))))))..))))------------------------))))).......)--))))))).)) ( -27.00, z-score = -2.82, R) >consensus ____UGUCUCCAUUUGGCAUUCAAAAACUGUGCGUCUGUGCGAUUUUGACGCAUAUUUGCCGGGCCCAAAACUACCUCACCCAAACU_UC___AUUCGCAUUGAUACUCACAUCGGUAC ............(((((((.........((((((((...........))))))))..)))))))....................................................... (-15.94 = -15.16 + -0.79)

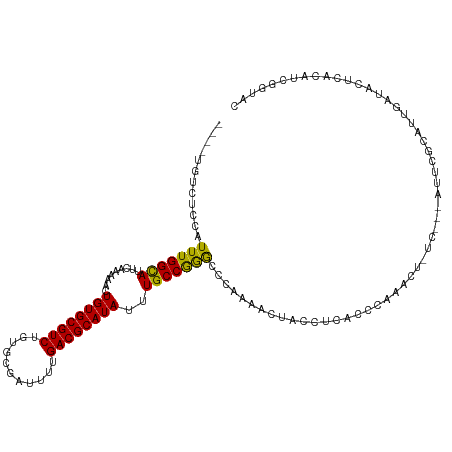

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:20:43 2011