
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 12,168,844 – 12,168,980 |
| Length | 136 |
| Max. P | 0.628704 |

| Location | 12,168,844 – 12,168,980 |
|---|---|
| Length | 136 |
| Sequences | 6 |
| Columns | 141 |
| Reading direction | reverse |
| Mean pairwise identity | 77.67 |
| Shannon entropy | 0.43014 |
| G+C content | 0.55646 |
| Mean single sequence MFE | -38.80 |
| Consensus MFE | -17.72 |
| Energy contribution | -19.28 |
| Covariance contribution | 1.56 |
| Combinations/Pair | 1.26 |
| Mean z-score | -1.93 |
| Structure conservation index | 0.46 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.29 |
| SVM RNA-class probability | 0.628704 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
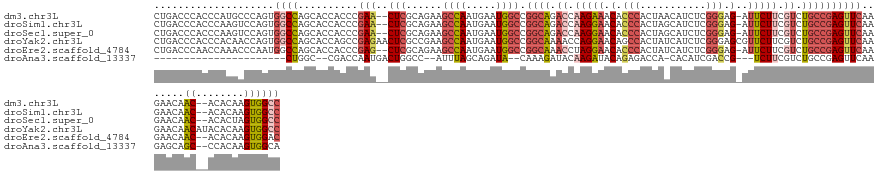
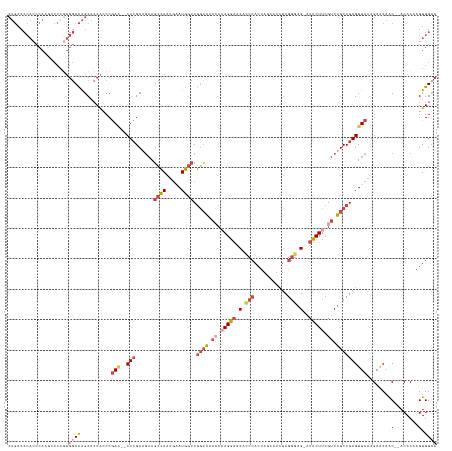
>dm3.chr3L 12168844 136 - 24543557 CUGACCCACCCAUGCCCAGUGGCCAGCACCACCCGAA--CUCGCAGAAGCCAAUGAAUGGCCGGCAGACCAAGAAACACCCACUAACAUCUCGGGAG-AUUCUUCGUCUGCCGAGUUCAAGAACAAC--ACACAAGUGGCC .....((((.........((((......))))..(((--(((......((((.....)))).(((((((.(((((.(.(((...........))).)-.))))).))))))))))))).........--......)))).. ( -45.60, z-score = -3.91, R) >droSim1.chr3L 11540158 136 - 22553184 CUGACCCACCCAAGUCCAGUGGCCAGCACCACCCGAA--CUCGCAGAAGCCAAUGAAUGGCCGGCAGACCAAGGAACACCCACUAGCAUCUCGGGAG-AUUCUUCGUCUGCCGAGUUCAAGAACAAC--ACACAAGUGGCC .....((((.........((((......))))..(((--(((......((((.....)))).(((((((.(((((.(.(((...........))).)-.))))).))))))))))))).........--......)))).. ( -45.10, z-score = -2.77, R) >droSec1.super_0 4353806 136 - 21120651 CUGACCCACCCAAGUCCAGUGGCCAGCACCACCCGAA--CUCGCAGAAGCCAAUGAAUGGCCGGCAGACCAAGGAACACCCACUAGCAUCUCGGGAG-AUUCUUCGUCUGCCGAGUUCAAGAACAAC--ACACUAGUGGCC .....((((.........((((......))))..(((--(((......((((.....)))).(((((((.(((((.(.(((...........))).)-.))))).))))))))))))).........--......)))).. ( -45.10, z-score = -2.73, R) >droYak2.chr3L 12216952 141 - 24197627 CUGACCCACCCACAACCAGUGGCCAGCACCAGCCGAGAACUCGCCGAAGCCAAUGAAUGGCCGGCAAAACCAGGAACAGCCACUAUCAUCUCGGGAGCGUUCUUCGUCUGCCGAGUUCAAGAACAACAUACACAAGUGGCC .........((((....((((((.....((....((....))((((..((((.....)))))))).......))....)))))).....(((((.((((.....)).)).)))))....................)))).. ( -40.60, z-score = -1.24, R) >droEre2.scaffold_4784 12157689 136 - 25762168 CUGACCCAACCAAACCCAAUGGCCAGCACCACCCGAG--CUCGCAGAAGCCAAUGAAUGGCCGGCAAACCUAGGAACACCCACUAUCAUCUCGGGAG-AUUCUUCGUCUGCCGAGUUCAAGAACAAC--ACACAAGUGGAC (((..(((...........))).)))..((((..(((--(((......((((.....)))).((((.((..((((.(.(((...........))).)-.))))..)).))))))))))..(.....)--......)))).. ( -33.20, z-score = -0.65, R) >droAna3.scaffold_13337 21816219 107 + 23293914 ----------------------CUGGC--CGACCAAUGACUGGCC--AUUUAGCAGAUA--CAAAGAUACAAGAUACAGAGACCA-CACAUCGACCG---UCUUCGUCUGCCGAGUUCAAGAGCAGC--CCACAAGUGGCA ----------------------..(((--((.........)))))--(((..((((((.--..(((((....(((..........-...)))....)---)))).))))))..))).........((--((....).))). ( -23.22, z-score = -0.30, R) >consensus CUGACCCACCCAAGCCCAGUGGCCAGCACCACCCGAA__CUCGCAGAAGCCAAUGAAUGGCCGGCAAACCAAGGAACACCCACUAGCAUCUCGGGAG_AUUCUUCGUCUGCCGAGUUCAAGAACAAC__ACACAAGUGGCC ....................(((((.......................((((.....))))(((((.((.(((((...(((...........)))....))))).)).))))).(((....)))............))))) (-17.72 = -19.28 + 1.56)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:20:25 2011