
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 12,011,704 – 12,011,801 |
| Length | 97 |
| Max. P | 0.891913 |

| Location | 12,011,704 – 12,011,801 |
|---|---|
| Length | 97 |
| Sequences | 11 |
| Columns | 101 |
| Reading direction | forward |
| Mean pairwise identity | 77.30 |
| Shannon entropy | 0.46831 |
| G+C content | 0.46764 |
| Mean single sequence MFE | -18.56 |
| Consensus MFE | -16.99 |
| Energy contribution | -17.40 |
| Covariance contribution | 0.41 |
| Combinations/Pair | 1.10 |
| Mean z-score | -0.82 |
| Structure conservation index | 0.92 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.10 |
| SVM RNA-class probability | 0.891913 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 12011704 97 + 24543557 ACCUGAGAUAUUGUUUCUCAACAUGCAAAUGCCGUGCGAUUUAU--UUGAGGAAGCGGCUAGCAUGC-CAAGACCCCAAACCCCAAACCCC-AAGCCCCCC ...(((((.......))))).(((((....(((((.(((.....--))).....)))))..))))).-.......................-......... ( -19.20, z-score = -1.47, R) >droSim1.chr3L 11381501 90 + 22553184 ACCUGAGAAAUUGUUUCUCAACAUGCAAAUGCCGUGCGAUUUAU--UUGAGGAAGCGGCUAGCAUGC-CAAAACCCCAAACCCCAAGACCCCC-------- ...((((((.....)))))).(((((....(((((.(((.....--))).....)))))..))))).-.........................-------- ( -19.60, z-score = -1.99, R) >droSec1.super_0 4197689 98 + 21120651 ACCUGAGAAAUUGUUUCUCAACAUGCAAAUGCUGUGCGAUUUAU--UUGAGGAAGCGGCUAGCAUGC-CAAAACCCCCAACCUCAAACCCCCAAGACCCCC ...((((((.....)))))).(((((....(((((.(((.....--))).....)))))..))))).-................................. ( -17.40, z-score = -0.83, R) >droYak2.chr3L 12057984 79 + 24197627 ACCUGAGAAAUUGUUUCUCAACAUGCAAAUGCCGUGCGAUUUAU--UUGAGGAAGCGGCUAGCAUGC-CAAAAUCCCAAACC------------------- ...((((((.....)))))).(((((....(((((.(((.....--))).....)))))..))))).-..............------------------- ( -19.60, z-score = -1.70, R) >droEre2.scaffold_4784 11999285 94 + 25762168 ACCUGAGAAAUUGUUUCUCAACAUGCAAAUGCCGUGCGAUUUAU--UUGAGGAAGCGGCUAGCAUGC-CAAAAC--CAAAACCAAAACCCCAAACUCCC-- ...((((((.....)))))).(((((....(((((.(((.....--))).....)))))..))))).-......--.......................-- ( -19.60, z-score = -1.76, R) >droAna3.scaffold_13337 11390828 96 - 23293914 ACCUGAGAAAUUGUUUCUCAACAUGCAAAUGCCGUGCGGUUUAU--UUGAGGAAGCGGCUCGCAUGC-CAAAAACCAAAGAGCCACAGAGCUAAAUCCA-- ...((((((.....)))))).....((((((((....)))..))--))).(((((((((((...((.-.......))..))))).....)))...))).-- ( -21.10, z-score = 0.25, R) >dp4.chrXR_group8 9050515 77 + 9212921 ACCUGAGAAAUUGUUUUUCAACAUGCAAAUGCCGUGCGAUUUAU--UUGAGGAAGCGGCUCGCAUGC-CAAACCAACACA--------------------- ...((((((.....)))))).(((((....(((((.(((.....--))).....)))))..))))).-............--------------------- ( -17.00, z-score = -0.37, R) >droPer1.super_50 399789 77 - 557249 ACCUGAGAAAUUGUUUUUCAACAUGCAAAUGCCGUGCGAUUUAU--UUGAGGAAGCGGCUCGCAUGC-CAAACCAACACA--------------------- ...((((((.....)))))).(((((....(((((.(((.....--))).....)))))..))))).-............--------------------- ( -17.00, z-score = -0.37, R) >droWil1.scaffold_180698 8517831 86 - 11422946 --CUGAGAAAUUGUUUCUCAACAUGCAAAUGCCGUGCGAUUUAU--UUGAGGAAGCGGCAGAGAUAAACGAGAGACGACAUGCCAACAAC----------- --........(((((((((..........((((((.(((.....--))).....)))))).........)))))))))............----------- ( -20.11, z-score = -1.66, R) >droMoj3.scaffold_6680 17624525 88 + 24764193 GCCUGAGAAAUUGUUUCUCCACAUGCAAAUGCCGUGCGAUUUAUAUUUGAGGAAGCGUCUCGCAUGC-CAAACGCCGAAAAGCCCCACA------------ ((..((((...((((((..((.(((.((((((...)).)))).))).))..))))))))))))....-.....((......))......------------ ( -14.20, z-score = 1.29, R) >droVir3.scaffold_13049 23011246 84 + 25233164 GCCUGAGAAAUUGUUUCUCCACAUGCAAAUGCCGUGCGAUUUAUAUUUGAGGAAGCGGCUCGCAUGC-CAAACC----AAAGCCCCACA------------ ((..(((((.....)))))..(((((....(((((....((((....))))...)))))..))))).-......----...))......------------ ( -19.30, z-score = -0.46, R) >consensus ACCUGAGAAAUUGUUUCUCAACAUGCAAAUGCCGUGCGAUUUAU__UUGAGGAAGCGGCUAGCAUGC_CAAAACCCCAAAACCCAAACA____________ ...((((((.....)))))).(((((....(((((....((((....))))...)))))..)))))................................... (-16.99 = -17.40 + 0.41)


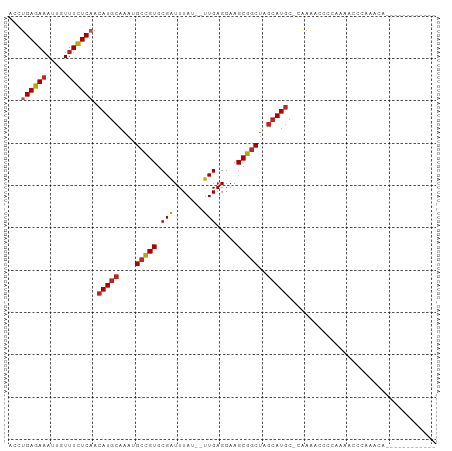
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:20:09 2011