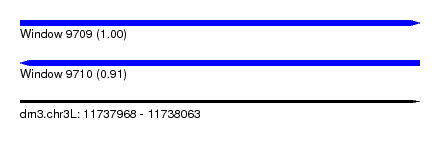

| Sequence ID | dm3.chr3L |
|---|---|
| Location | 11,737,968 – 11,738,063 |
| Length | 95 |
| Max. P | 0.996005 |

| Location | 11,737,968 – 11,738,063 |
|---|---|
| Length | 95 |
| Sequences | 6 |
| Columns | 103 |
| Reading direction | forward |
| Mean pairwise identity | 60.15 |
| Shannon entropy | 0.75350 |
| G+C content | 0.38587 |
| Mean single sequence MFE | -19.80 |
| Consensus MFE | -7.97 |
| Energy contribution | -8.67 |
| Covariance contribution | 0.70 |
| Combinations/Pair | 1.41 |
| Mean z-score | -2.13 |
| Structure conservation index | 0.40 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.87 |
| SVM RNA-class probability | 0.996005 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 11737968 95 + 24543557 CUUUAGAAAAGGAAAAGUUCCAAUGUUUUUUUCCCAUCCCACUUUUGGCCUGCCAGAGAUAUAACUAUCUUAUGAUAGGAACUUUUUCCCACUUU-------- ..........(((((((((((.....................((((((....)))))).......((((....))))))))))))))).......-------- ( -21.40, z-score = -2.42, R) >droSim1.chrU 245641 94 - 15797150 CUUUAAAGCAGGAAAAGUUCCAAUGUUUUGC-CCCAUCCCACUUUUGACCUCCCAGAGAUAUUCCUAUCUUAUGAUAGGAACUUUUCCCCACUUU-------- ....((((..(((((((((((((((((((..-....((........)).......))))))))..((((....)))))))))))))))...))))-------- ( -21.02, z-score = -3.18, R) >droSec1.super_0 3923549 89 + 21120651 CUUUAAAGCAGGAAA-GUUCCAAUGUUUUUC-CUCAUCCCAUUUUUGGC----CAGAGAUAUUCCUAUCUUAUGAUUGGAACUUUUCCCCACUUU-------- ..........(((((-((((((((.......-......(((....))).----..((((((....))))))...)))))))))))))........-------- ( -21.60, z-score = -2.94, R) >droYak2.chr3L 11766539 103 + 24197627 GUUUAAAGACGGAAAAGUUCCAACGUUUUUGCCCAAUCCCACUUUCGGCCUCCCAGAGAUAUUUCUAUCUAAUAAAAGGAACUUUGCCAACGUUCGCCACUUU (((....)))((.((((((((.........(((.((.......)).))).......(((((....))))).......)))))))).))............... ( -19.10, z-score = -1.17, R) >droEre2.scaffold_4784 11716531 95 + 25762168 GCUUAAAGGAGGAAAAGUUUCAGUGUUUUUCCCCAAUC--------GGCCUCCCAGAGAUAUUUCUGUCUAAUGAAAGGAACUUUACCAUCGUUCGCCACUUU .......((.(((((((........)))))))))....--------(((....(((((....)))))...........((((.........)))))))..... ( -18.70, z-score = 0.00, R) >droAna3.scaffold_13337 5993678 96 + 23293914 UUUUACGUAAUAUUACGUAUACGCAAUGUUUGCUGAAU-------UGAACUUUCUUUUCAAGCCCUAACAUUCCCUGUACCCUUUUCUCUCAGUGAAGUAUUU (((((((((....)))(((((.(.((((((.((((((.-------.((....))..))).)))...)))))).).)))))............))))))..... ( -17.00, z-score = -3.07, R) >consensus CUUUAAAGAAGGAAAAGUUCCAAUGUUUUUC_CCCAUCCCACUUUUGGCCUCCCAGAGAUAUUCCUAUCUUAUGAUAGGAACUUUUCCCCACUUU________ ..........((.((((((((.........................((....))..(((((....))))).......)))))))).))............... ( -7.97 = -8.67 + 0.70)


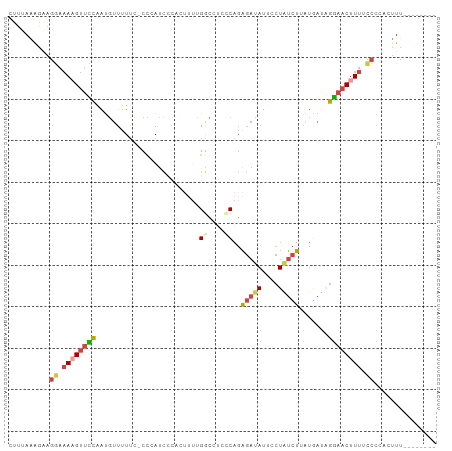
| Location | 11,737,968 – 11,738,063 |
|---|---|
| Length | 95 |
| Sequences | 6 |
| Columns | 103 |
| Reading direction | reverse |
| Mean pairwise identity | 60.15 |
| Shannon entropy | 0.75350 |
| G+C content | 0.38587 |
| Mean single sequence MFE | -23.04 |
| Consensus MFE | -9.02 |
| Energy contribution | -9.08 |
| Covariance contribution | 0.06 |
| Combinations/Pair | 1.65 |
| Mean z-score | -1.32 |
| Structure conservation index | 0.39 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.21 |
| SVM RNA-class probability | 0.910187 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 11737968 95 - 24543557 --------AAAGUGGGAAAAAGUUCCUAUCAUAAGAUAGUUAUAUCUCUGGCAGGCCAAAAGUGGGAUGGGAAAAAAACAUUGGAACUUUUCCUUUUCUAAAG --------..((.(((((((.(((((((((....))))....(((((((((....))).....)))))).............))))))))))))...)).... ( -22.30, z-score = -1.12, R) >droSim1.chrU 245641 94 + 15797150 --------AAAGUGGGGAAAAGUUCCUAUCAUAAGAUAGGAAUAUCUCUGGGAGGUCAAAAGUGGGAUGGG-GCAAAACAUUGGAACUUUUCCUGCUUUAAAG --------.....(((((...(((((((((....))))))))).))))).........((((..(((..((-.(((....)))...))..)))..)))).... ( -27.20, z-score = -2.38, R) >droSec1.super_0 3923549 89 - 21120651 --------AAAGUGGGGAAAAGUUCCAAUCAUAAGAUAGGAAUAUCUCUG----GCCAAAAAUGGGAUGAG-GAAAAACAUUGGAAC-UUUCCUGCUUUAAAG --------((((..((((..((((((((((((.(((((....)))))...----.(((....))).))))(-......).)))))))-)))))..)))).... ( -25.50, z-score = -2.90, R) >droYak2.chr3L 11766539 103 - 24197627 AAAGUGGCGAACGUUGGCAAAGUUCCUUUUAUUAGAUAGAAAUAUCUCUGGGAGGCCGAAAGUGGGAUUGGGCAAAAACGUUGGAACUUUUCCGUCUUUAAAC (((((..(.((((((.((..((((((((((...(((((....)))))...))))..(....).))))))..))...)))))).).)))))............. ( -26.40, z-score = -0.96, R) >droEre2.scaffold_4784 11716531 95 - 25762168 AAAGUGGCGAACGAUGGUAAAGUUCCUUUCAUUAGACAGAAAUAUCUCUGGGAGGCC--------GAUUGGGGAAAAACACUGAAACUUUUCCUCCUUUAAGC ((((((((((((.........))))((..((..(((.(....).))).))..)))))--------)...((((((((..........)))))))))))).... ( -21.20, z-score = 0.11, R) >droAna3.scaffold_13337 5993678 96 - 23293914 AAAUACUUCACUGAGAGAAAAGGGUACAGGGAAUGUUAGGGCUUGAAAAGAAAGUUCA-------AUUCAGCAAACAUUGCGUAUACGUAAUAUUACGUAAAA ...(((....(((.............)))(.((((((...(((.(((..((....)).-------.)))))).)))))).))))((((((....))))))... ( -15.62, z-score = -0.65, R) >consensus ________AAAGGGGGGAAAAGUUCCUAUCAUAAGAUAGGAAUAUCUCUGGGAGGCCAAAAGUGGGAUGGG_GAAAAACAUUGGAACUUUUCCUUCUUUAAAG ........((((((.((.((((((((.......(((((....)))))..((....)).........................)))))))).)))))))).... ( -9.02 = -9.08 + 0.06)



Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:19:33 2011