
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 4,864,169 – 4,864,233 |
| Length | 64 |
| Max. P | 0.540157 |

| Location | 4,864,169 – 4,864,233 |
|---|---|
| Length | 64 |
| Sequences | 9 |
| Columns | 72 |
| Reading direction | reverse |
| Mean pairwise identity | 82.38 |
| Shannon entropy | 0.34986 |
| G+C content | 0.60280 |
| Mean single sequence MFE | -27.08 |
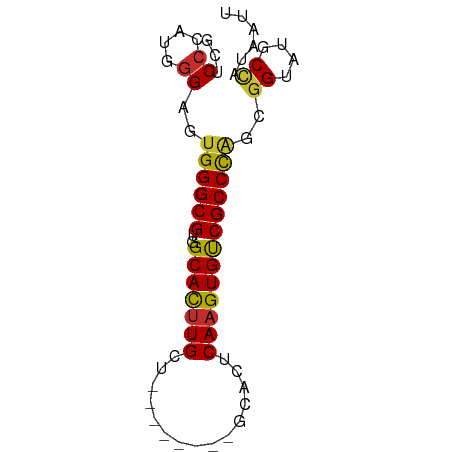
| Consensus MFE | -17.08 |
| Energy contribution | -16.98 |
| Covariance contribution | -0.09 |
| Combinations/Pair | 1.30 |
| Mean z-score | -1.56 |
| Structure conservation index | 0.63 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.09 |
| SVM RNA-class probability | 0.540157 |
| Prediction | RNA |
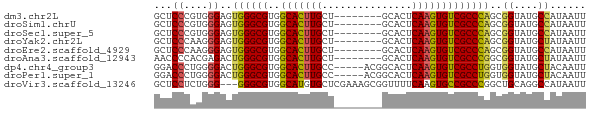
Download alignment: ClustalW | MAF
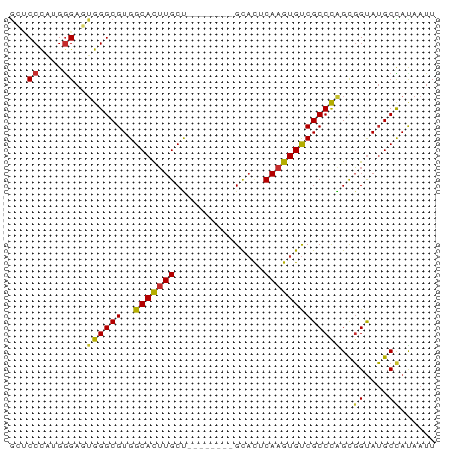
>dm3.chr2L 4864169 64 - 23011544 GCUCCCGUGGGAGUGGGCGUGGCACUUGCU--------GCACUCAAGUGUCGCCCAGCGGUAUGCCAUAAUU (((((....)))))(((((..(((((((..--------.....))))))))))))...((....))...... ( -27.00, z-score = -1.55, R) >droSim1.chrU 15295428 64 - 15797150 GCUCCCGUGGGAGUGGGCGUGGCACUUGCU--------GCACUCAAGUGUCGCCCAGCGGUAUGCCAUAAUU (((((....)))))(((((..(((((((..--------.....))))))))))))...((....))...... ( -27.00, z-score = -1.55, R) >droSec1.super_5 2959300 64 - 5866729 GCUCCCGUGGGAGUGGGCGUGGCACUUGCU--------GCACUCAAGUGUCGCCCAGCGGUAUGCCAUAAUU (((((....)))))(((((..(((((((..--------.....))))))))))))...((....))...... ( -27.00, z-score = -1.55, R) >droYak2.chr2L 4874576 64 - 22324452 GCUCCCAAGGGAGUGGGCGUGGCACUUGCU--------GCACUCAAGUGUCGCCCAGCGGUAUGCUAUAAUU (((((....)))))(((((..(((((((..--------.....))))))))))))(((.....)))...... ( -26.60, z-score = -1.96, R) >droEre2.scaffold_4929 4945785 64 - 26641161 GCUCCCAAGGGAGUGGGCGUGGCACUUGCU--------GCACUCAAGUGUCGCCCAGCGGUAUGCCAUAAUU (((((....)))))(((((..(((((((..--------.....))))))))))))...((....))...... ( -27.00, z-score = -1.87, R) >droAna3.scaffold_12943 2603769 64 - 5039921 AACCCCACGAGACUGGGCGUGGCACUUGCU--------GCACUCAAGUGUCGCCCGGCGGUAUGCUAUAAUU .(((((.(((.((((((.(..((....)).--------.).))).))).)))...)).)))........... ( -21.90, z-score = -0.98, R) >dp4.chr4_group3 6109705 67 + 11692001 GGACCCUGGGGACUGGGCGUGGCACUUGCC-----ACGGCACUCAAGUGUCGCCUGGUGGUAUGCUACAAUU ...((....))((..((((((((....)))-----))(((((....))))))))..)).............. ( -28.90, z-score = -1.77, R) >droPer1.super_1 3208282 67 + 10282868 GGACCCUGGGGACUGGGCGUGGCACUUGCC-----ACGGCACUCAAGUGUCGCCUGGUGGUAUGCUACAAUU ...((....))((..((((((((....)))-----))(((((....))))))))..)).............. ( -28.90, z-score = -1.77, R) >droVir3.scaffold_13246 2269961 69 - 2672878 GCUCCUCUGGG---GGGCGUGGCAUGUGCUCGAAAGCGGUUUUCAAGUGCCGCCCGGCUGCAGGCCAUAAUU (((((.....)---))))(((((.(((((.((...(((((........))))).)))).))).))))).... ( -29.40, z-score = -1.02, R) >consensus GCUCCCAUGGGAGUGGGCGUGGCACUUGCU________GCACUCAAGUGUCGCCCAGCGGUAUGCCAUAAUU ((.((....)).)).(((((((((((((...............))))))))(((....))))))))...... (-17.08 = -16.98 + -0.09)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:17:38 2011