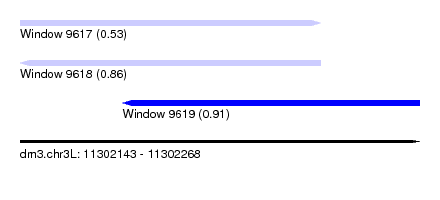

| Sequence ID | dm3.chr3L |
|---|---|
| Location | 11,302,143 – 11,302,268 |
| Length | 125 |
| Max. P | 0.907926 |

| Location | 11,302,143 – 11,302,237 |
|---|---|
| Length | 94 |
| Sequences | 7 |
| Columns | 99 |
| Reading direction | forward |
| Mean pairwise identity | 73.36 |
| Shannon entropy | 0.52646 |
| G+C content | 0.42661 |
| Mean single sequence MFE | -20.04 |
| Consensus MFE | -6.93 |
| Energy contribution | -7.91 |
| Covariance contribution | 0.99 |
| Combinations/Pair | 1.28 |
| Mean z-score | -1.92 |
| Structure conservation index | 0.35 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.08 |
| SVM RNA-class probability | 0.532453 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 11302143 94 + 24543557 GAUGACGGCACACAAGUUGACCGCAACCAAUCC-GAAUUAGUUUCGGAUUAUCUCCUACACAAUUGCACAAGAUAAUUGAGGUAGCAGGGCAUAU---- .(((.((((......)))).((((.((((((((-(((.....))))))))..........(((((((....).)))))).))).)).)).)))..---- ( -23.00, z-score = -2.16, R) >droSim1.chr3L 10691074 84 + 22553184 GAUGACGGCACACAACUUGACCGCAACCAAUCC-GAAUUAGUUUCGGAUUAUCUCCUACACAAUUGCACAAGAUAAUUGAGGUAG-------------- ..((.(((((.......)).)))))((((((((-(((.....))))))))..........(((((((....).)))))).)))..-------------- ( -18.90, z-score = -2.58, R) >droSec1.super_0 3515135 90 + 21120651 GAUGACGGCACACAACUUGACCGCAACCAAUCC-GAAUUAGUUUCGGAUUAUCUCCUACACAAUUGCACAAGAUAAUUGAGGUAGCAGGAC-------- ..((.(((((.......)).)))))...(((((-(((.....)))))))).(((.((((.(((((((....).))))))..)))).)))..-------- ( -21.60, z-score = -2.64, R) >droYak2.chr3L 11337469 96 + 24197627 GAUGACGGCACACAACUUGACCGCAACCAAUCC-GAAUUAGUUUCGGAUUAUCUCUUACACAAUUGCACCAGAUAAUUGAGGUAGCAGGGCAUACAU-- .(((...((.((.....)).((((.((((((((-(((.....))))))))..........(((((((....).)))))).))).)).))))...)))-- ( -21.40, z-score = -1.92, R) >droEre2.scaffold_4784 11305945 92 + 25762168 GAUGACGGCACACAACUUGACCGCAACCAAUCC-GAAUUAGUUUCGGAUUAUCUCUUACACAAUUGCACCAGAUAAUUGAGGUAGCAGGGCAU------ .......((.((.....)).((((.((((((((-(((.....))))))))..........(((((((....).)))))).))).)).))))..------ ( -20.90, z-score = -1.73, R) >droAna3.scaffold_13337 5586143 91 + 23293914 GAUGACGGCACACAACUCGACCGCAACAAAUCCCGAAUCAGUUUCGGAUUAUCUUAUACACAAUUACGCAAGAUAAUUGAGGUAGAAAAGG-------- (((((((((.........).))).........(((((.....))))).)))))...(((.(((((((....).))))))..))).......-------- ( -17.20, z-score = -1.66, R) >droVir3.scaffold_13049 1592962 93 - 25233164 ---GAGAGAGUGGAAAUUAAUUUCAUUCAAUGGCGAG--GACGCCAAAUGGACUUA-AUGUUGUUAAAUAGCUUAGAUGAUUCAGCACAAGAAAAACGG ---(((.((((((((.....))))))))..(((((..--..)))))......))).-.((((........(((..((....)))))........)))). ( -17.29, z-score = -0.77, R) >consensus GAUGACGGCACACAACUUGACCGCAACCAAUCC_GAAUUAGUUUCGGAUUAUCUCCUACACAAUUGCACAAGAUAAUUGAGGUAGCAGGGCA_______ ..((.(((((.......)).)))))((((((((.(((.....))))))))..........(((((((....).)))))).)))................ ( -6.93 = -7.91 + 0.99)

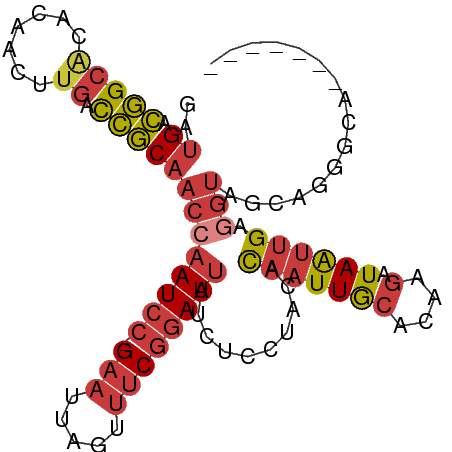
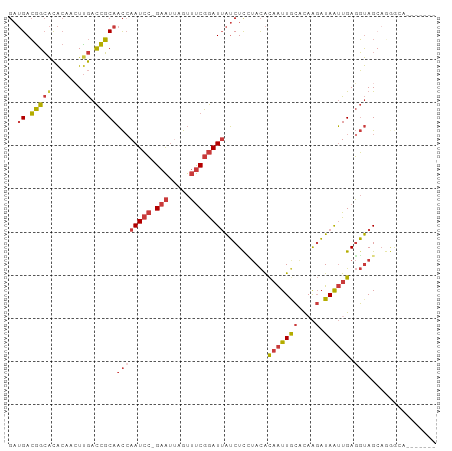
| Location | 11,302,143 – 11,302,237 |
|---|---|
| Length | 94 |
| Sequences | 7 |
| Columns | 99 |
| Reading direction | reverse |
| Mean pairwise identity | 73.36 |
| Shannon entropy | 0.52646 |
| G+C content | 0.42661 |
| Mean single sequence MFE | -22.03 |
| Consensus MFE | -11.17 |
| Energy contribution | -12.26 |
| Covariance contribution | 1.09 |
| Combinations/Pair | 1.41 |
| Mean z-score | -1.55 |
| Structure conservation index | 0.51 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.95 |
| SVM RNA-class probability | 0.859753 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 11302143 94 - 24543557 ----AUAUGCCCUGCUACCUCAAUUAUCUUGUGCAAUUGUGUAGGAGAUAAUCCGAAACUAAUUC-GGAUUGGUUGCGGUCAACUUGUGUGCCGUCAUC ----.......((((.(((....(((((((.((((....)))).)))))))((((((.....)))-)))..))).)))).................... ( -25.50, z-score = -1.98, R) >droSim1.chr3L 10691074 84 - 22553184 --------------CUACCUCAAUUAUCUUGUGCAAUUGUGUAGGAGAUAAUCCGAAACUAAUUC-GGAUUGGUUGCGGUCAAGUUGUGUGCCGUCAUC --------------..(((....(((((((.((((....)))).)))))))((((((.....)))-)))..))).(((((((.....)).))))).... ( -23.60, z-score = -2.40, R) >droSec1.super_0 3515135 90 - 21120651 --------GUCCUGCUACCUCAAUUAUCUUGUGCAAUUGUGUAGGAGAUAAUCCGAAACUAAUUC-GGAUUGGUUGCGGUCAAGUUGUGUGCCGUCAUC --------...((((.(((....(((((((.((((....)))).)))))))((((((.....)))-)))..))).)))).................... ( -25.50, z-score = -2.37, R) >droYak2.chr3L 11337469 96 - 24197627 --AUGUAUGCCCUGCUACCUCAAUUAUCUGGUGCAAUUGUGUAAGAGAUAAUCCGAAACUAAUUC-GGAUUGGUUGCGGUCAAGUUGUGUGCCGUCAUC --..((((((...((((((.(((((((((..((((....))))..))))((((((((.....)))-)))))))))).)))..))).))))))....... ( -26.10, z-score = -1.90, R) >droEre2.scaffold_4784 11305945 92 - 25762168 ------AUGCCCUGCUACCUCAAUUAUCUGGUGCAAUUGUGUAAGAGAUAAUCCGAAACUAAUUC-GGAUUGGUUGCGGUCAAGUUGUGUGCCGUCAUC ------..((((.((((((.(((((((((..((((....))))..))))((((((((.....)))-)))))))))).)))..))).).).))....... ( -23.00, z-score = -1.13, R) >droAna3.scaffold_13337 5586143 91 - 23293914 --------CCUUUUCUACCUCAAUUAUCUUGCGUAAUUGUGUAUAAGAUAAUCCGAAACUGAUUCGGGAUUUGUUGCGGUCGAGUUGUGUGCCGUCAUC --------.(..(((.(((.(((((((((((..((....))..))))))).((((((.....))))))....)))).))).)))..)............ ( -18.50, z-score = -0.50, R) >droVir3.scaffold_13049 1592962 93 + 25233164 CCGUUUUUCUUGUGCUGAAUCAUCUAAGCUAUUUAACAACAU-UAAGUCCAUUUGGCGUC--CUCGCCAUUGAAUGAAAUUAAUUUCCACUCUCUC--- ..(((........(((..........)))........)))((-(((...(((((((((..--..)))))...))))...)))))............--- ( -11.99, z-score = -0.60, R) >consensus _______UGCCCUGCUACCUCAAUUAUCUUGUGCAAUUGUGUAGGAGAUAAUCCGAAACUAAUUC_GGAUUGGUUGCGGUCAAGUUGUGUGCCGUCAUC ......................((((((((.((((....)))).))))))))(((.(((((((.....))))))).))).................... (-11.17 = -12.26 + 1.09)

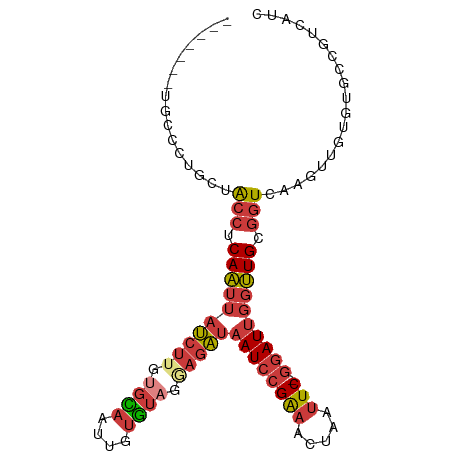
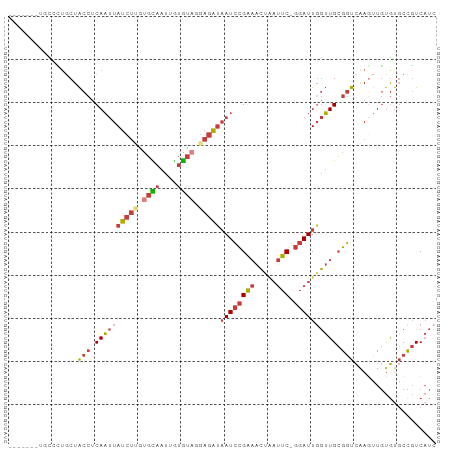
| Location | 11,302,175 – 11,302,268 |
|---|---|
| Length | 93 |
| Sequences | 6 |
| Columns | 99 |
| Reading direction | reverse |
| Mean pairwise identity | 70.97 |
| Shannon entropy | 0.54310 |
| G+C content | 0.36809 |
| Mean single sequence MFE | -16.78 |
| Consensus MFE | -8.69 |
| Energy contribution | -8.97 |
| Covariance contribution | 0.28 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.55 |
| Structure conservation index | 0.52 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.20 |
| SVM RNA-class probability | 0.907926 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
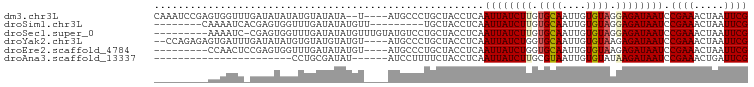
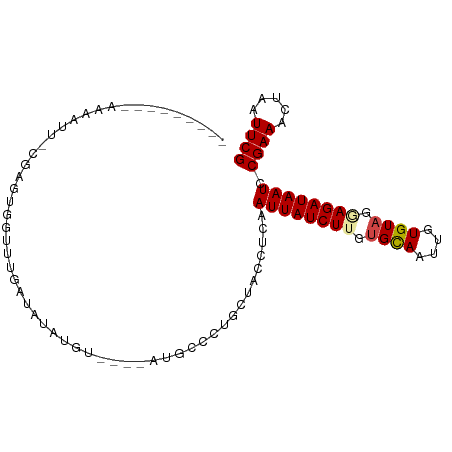
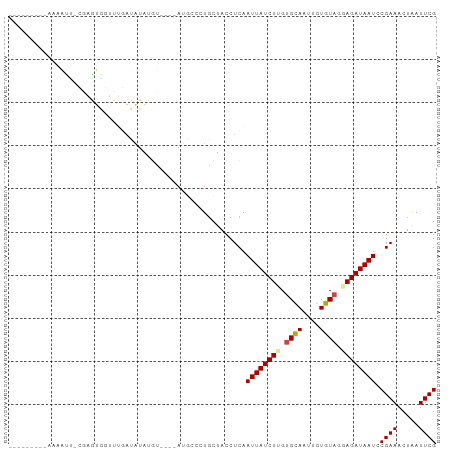
>dm3.chr3L 11302175 93 - 24543557 CAAAUCCGAGUGGUUUGAUAUAUAUGUAUAUA--U----AUGCCCUGCUACCUCAAUUAUCUUGUGCAAUUGUGUAGGAGAUAAUCCGAAACUAAUUCG .......(((((((..(.(((((((....)))--)----))).)..)))).))).((((((((.((((....)))).)))))))).((((.....)))) ( -21.20, z-score = -2.27, R) >droSim1.chr3L 10691106 82 - 22553184 --------CAAAAUCACGAGUGGUUUGAUAUAUGUU---------UGCUACCUCAAUUAUCUUGUGCAAUUGUGUAGGAGAUAAUCCGAAACUAAUUCG --------.........(((((((..(((....)))---------.)))).))).((((((((.((((....)))).)))))))).((((.....)))) ( -16.90, z-score = -1.41, R) >droSec1.super_0 3515167 89 - 21120651 ---------AAAAUC-CGAGUGGUUUGAUAUAUGUUUGUAUGUCCUGCUACCUCAAUUAUCUUGUGCAAUUGUGUAGGAGAUAAUCCGAAACUAAUUCG ---------......-.(((((((..(((((((....)))))))..)))).))).((((((((.((((....)))).)))))))).((((.....)))) ( -24.50, z-score = -4.14, R) >droYak2.chr3L 11337501 93 - 24197627 --CCAGAGAGUGAUUUGAUAUAUGUGUAUGUAUGU----AUGCCCUGCUACCUCAAUUAUCUGGUGCAAUUGUGUAAGAGAUAAUCCGAAACUAAUUCG --...(((.(((.(..(.((((((((....)))))----))).)..).)))))).(((((((..((((....))))..))))))).((((.....)))) ( -15.70, z-score = -0.17, R) >droEre2.scaffold_4784 11305977 86 - 25762168 ---------CCAACUCCGAGUGGUUUGAUAUAUGU----AUGCCCUGCUACCUCAAUUAUCUGGUGCAAUUGUGUAAGAGAUAAUCCGAAACUAAUUCG ---------.......((((((((((.......((----(.((...)))))....(((((((..((((....))))..)))))))...))))).))))) ( -14.90, z-score = -0.55, R) >droAna3.scaffold_13337 5586176 70 - 23293914 -----------------------CCUGCGAUAU------AUCCUUUUCUACCUCAAUUAUCUUGCGUAAUUGUGUAUAAGAUAAUCCGAAACUGAUUCG -----------------------.....(((.(------(((.((...(((..(((((((.....))))))).))).)))))))))((((.....)))) ( -7.50, z-score = -0.75, R) >consensus _________AAAAUU_CGAGUGGUUUGAUAUAUGU____AUGCCCUGCUACCUCAAUUAUCUUGUGCAAUUGUGUAGGAGAUAAUCCGAAACUAAUUCG .......................................................((((((((.((((....)))).)))))))).((((.....)))) ( -8.69 = -8.97 + 0.28)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:18:18 2011