| Sequence ID | dm3.chr3L |
|---|---|
| Location | 11,291,098 – 11,291,161 |
| Length | 63 |
| Max. P | 0.989162 |

| Location | 11,291,098 – 11,291,161 |
|---|---|
| Length | 63 |
| Sequences | 5 |
| Columns | 63 |
| Reading direction | reverse |
| Mean pairwise identity | 96.98 |
| Shannon entropy | 0.05258 |
| G+C content | 0.50159 |
| Mean single sequence MFE | -21.64 |
| Consensus MFE | -20.62 |
| Energy contribution | -21.02 |
| Covariance contribution | 0.40 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.71 |
| Structure conservation index | 0.95 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.35 |
| SVM RNA-class probability | 0.989162 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
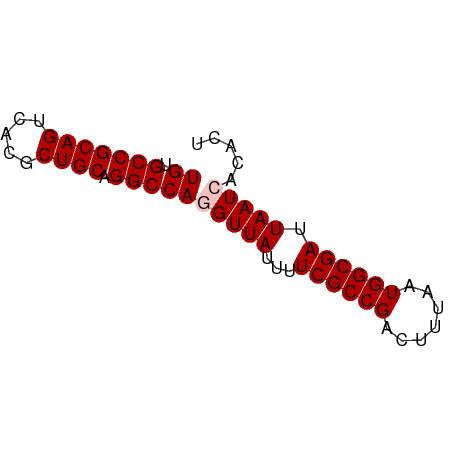
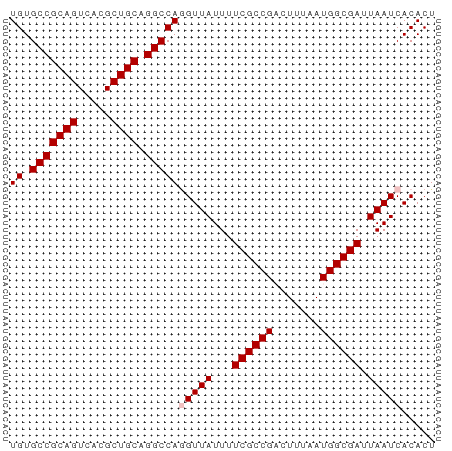
>dm3.chr3L 11291098 63 - 24543557 UGUGCCGCAGUCACGCUGCAGGCCAGGUUAUUUUCGCCGACUUUAAUGGCGAUUAAUCACACU ((.(((((((.....)))).)))))(((((...((((((.......)))))).)))))..... ( -22.50, z-score = -3.00, R) >droEre2.scaffold_4784 11294629 63 - 25762168 UGUGCCGCAGUCACGCUGCAGGCCAAGUUAUUUUCGCCGCUUUUAAUGGCGAUUAAUCACACU ((.(((((((.....)))).))))).((.(((.((((((.......))))))..))).))... ( -20.20, z-score = -2.28, R) >droYak2.chr3L 11325635 63 - 24197627 UGUGCCGCAGUCACGCUGCAGGCCAUGUUAUUUUCGCCGCUUUUAAUGGCGAUUAAUCACACU ((.(((((((.....)))).)))))(((.(((.((((((.......))))))..))).))).. ( -20.50, z-score = -2.28, R) >droSec1.super_0 3504163 63 - 21120651 UGUGCCGCAGUCACGCUGCAGGCCAGGUUAUUUUCGCCGACUUUAAUGGCGAUUAAUCACACU ((.(((((((.....)))).)))))(((((...((((((.......)))))).)))))..... ( -22.50, z-score = -3.00, R) >droSim1.chr3L 10679611 63 - 22553184 UGUGCCGCAGUCACGCUGCAGGCCAGGUUAUUUUCGCCGACUUUAAUGGCGAUUAAUCACACU ((.(((((((.....)))).)))))(((((...((((((.......)))))).)))))..... ( -22.50, z-score = -3.00, R) >consensus UGUGCCGCAGUCACGCUGCAGGCCAGGUUAUUUUCGCCGACUUUAAUGGCGAUUAAUCACACU ((.(((((((.....)))).)))))(((((...((((((.......)))))).)))))..... (-20.62 = -21.02 + 0.40)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:18:12 2011