
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 11,265,575 – 11,265,682 |
| Length | 107 |
| Max. P | 0.583317 |

| Location | 11,265,575 – 11,265,682 |
|---|---|
| Length | 107 |
| Sequences | 8 |
| Columns | 109 |
| Reading direction | reverse |
| Mean pairwise identity | 83.89 |
| Shannon entropy | 0.30742 |
| G+C content | 0.51055 |
| Mean single sequence MFE | -24.77 |
| Consensus MFE | -15.24 |
| Energy contribution | -14.94 |
| Covariance contribution | -0.30 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.85 |
| Structure conservation index | 0.62 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.18 |
| SVM RNA-class probability | 0.583317 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
>dm3.chr3L 11265575 107 - 24543557 -GUUCGCCAUGCGCUUUGCAUGCCAAGUGUAUCCACAUCCAGACCUGUGAUUCACUGAUAACACGUGCCAAGUGCACUCCACCUGACCCAAACUCAGAACCAAAACCA- -((((....(((((((.(((((...((((.(((.(((........)))))).)))).......))))).))))))).......(((.......)))))))........- ( -24.70, z-score = -2.19, R) >droSim1.chr3L 10654438 107 - 22553184 -GUUCGCCAUGCGCUUUGCAUGCCAAGUGUAUCCACAUCCAGACCUGUGAUUCACUGAUAACACGUGCCAAGUGCACUCCGCCUGACCCAAACCCACAACCAAAACCA- -(((.((..(((((((.(((((...((((.(((.(((........)))))).)))).......))))).)))))))....))..))).....................- ( -22.60, z-score = -1.63, R) >droSec1.super_0 3480409 107 - 21120651 -GUUCGCCAUGCGCUUUGCAUGCCAAGUGUAUCCACAUCCAGACCUGUGAUUCACUGAUAACACGUGCCAAGUGCACUCCACCUGACCCAAACCCACAACCAAAACCA- -........(((((((.(((((...((((.(((.(((........)))))).)))).......))))).)))))))................................- ( -21.90, z-score = -1.64, R) >droYak2.chr3L 11300086 101 - 24197627 -GUCCGCCAUGCGCUUUGCAUGCCAAGUAUAUCCACAUCCAGACCUCUGAUUCACUGAUAACACGUGCCAAGUGCACUCCACCUGACCCAAGCCAAAAACCA------- -(((.....(((((((.(((((....((.((((......(((....))).......)))))).))))).)))))))........)))...............------- ( -18.24, z-score = -1.28, R) >droEre2.scaffold_4784 11269760 100 - 25762168 -GUGCGCCAUGCGCUUUGCAUGCCAAGUAUAUCCACAUCCAGACCUCUGAUUCACUGAUAACACGUGCCAAGUGCACUACACCUGACCCAAAC-CAAAACCA------- -(((.....(((((((.(((((....((.((((......(((....))).......)))))).))))).)))))))...)))...........-........------- ( -18.92, z-score = -1.25, R) >droAna3.scaffold_13337 5553024 100 - 23293914 -GCCCUACAUGCGCUUUGCAUGCCAAGUAUAUCCACAUCCAGACCUCUGAUUCAUUGAUAACACGUGCCAAGUGCACUACACCUGACCC-AGCCCAAAACCA------- -........(((((((.(((((....((.((((......(((....))).......)))))).))))).))))))).............-............------- ( -18.12, z-score = -1.68, R) >droPer1.super_20 1480903 109 + 1872136 GGCUCUCCAUGCGCUUUGCAUGCUAAGUGUAUCCAUAUCCAGACCUCUGAUUCAUUGAUAACACGUGCUGAGUGCACUACACCUGCCUGGAAGACAGGCAGACAACCCA ((.......(((((((.((((.....(((((((.(((..(((....)))..).)).))).)))))))).)))))))......(((((((.....))))))).....)). ( -33.80, z-score = -2.20, R) >dp4.chrXR_group6 2875267 109 + 13314419 GGCUCUCCAUGCGCUUUGCAUGCUAAGUGUAUCCAUAUCCAGACCUCUGAUUCAUGGAUAACACGUGCUGGGUGCACUACACCUGCCUGGAAGACAGGCAGACAACCCA ((.......((((((..((((.....(((((((((((..(((....)))..).)))))).))))))))..))))))......(((((((.....))))))).....)). ( -39.90, z-score = -2.90, R) >consensus _GUUCGCCAUGCGCUUUGCAUGCCAAGUGUAUCCACAUCCAGACCUCUGAUUCACUGAUAACACGUGCCAAGUGCACUACACCUGACCCAAACCCAAAACCA_AACC__ .........(((((((.(((((....((.((((......((......)).......)))))).))))).)))))))................................. (-15.24 = -14.94 + -0.30)
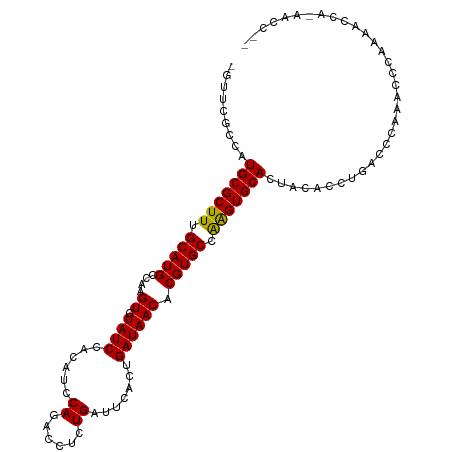


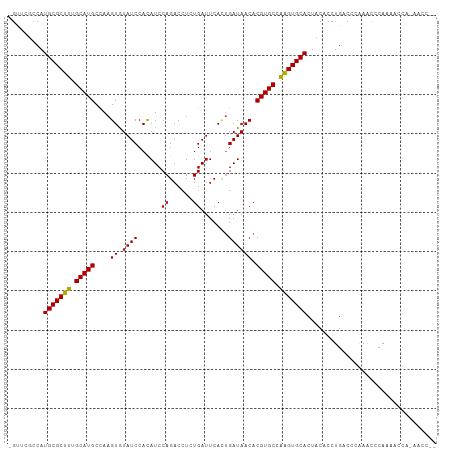
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:18:10 2011