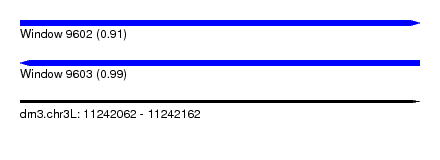

| Sequence ID | dm3.chr3L |
|---|---|
| Location | 11,242,062 – 11,242,162 |
| Length | 100 |
| Max. P | 0.990345 |

| Location | 11,242,062 – 11,242,162 |
|---|---|
| Length | 100 |
| Sequences | 4 |
| Columns | 100 |
| Reading direction | forward |
| Mean pairwise identity | 63.57 |
| Shannon entropy | 0.55743 |
| G+C content | 0.37051 |
| Mean single sequence MFE | -19.82 |
| Consensus MFE | -6.96 |
| Energy contribution | -10.03 |
| Covariance contribution | 3.06 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.36 |
| Structure conservation index | 0.35 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.24 |
| SVM RNA-class probability | 0.914909 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
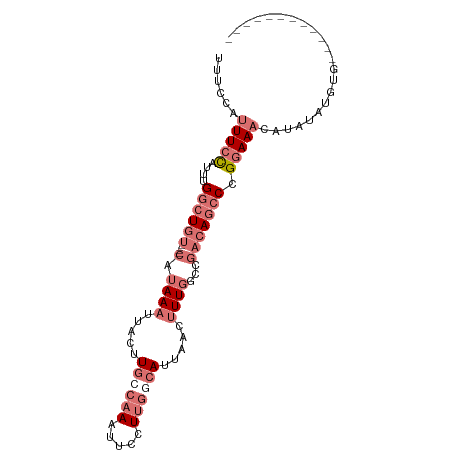
>dm3.chr3L 11242062 100 + 24543557 UUUCCAUUUCCAUUUGGCUGUCUGUCAUAAAUUACUUGCCAAAUUCCUUGGCAUUAACUUUGGCCGACAGCCCGGAAACAUAACUGUGUACAUACUUACA .......((((....(((((((.((((....(((..((((((.....)))))).)))...)))).))))))).))))((((....))))........... ( -28.80, z-score = -4.51, R) >droSim1.chr3L 10636345 84 + 22553184 UUUCCAUUUCCAUUUGGCUGU----CAUAAAUUACUUGCCAAAUUCCUUGGCAUUAACUUUGGCCGACAGCCCGGAAACAUAUCUGUG------------ ......(((((....((((((----(.((((.....((((((.....)))))).....))))...))))))).)))))..........------------ ( -24.00, z-score = -3.77, R) >droEre2.scaffold_4784 11252928 80 + 25762168 UUCCCAUUUCCAUUUGGCUGU----CAUAAAUUACUUGGCAAAUUCCUUGGCAUUAACUUUGGCCGACAGCCCGGAAACAUAUA---------------- ......(((((....((((((----(...........((......))..(((..........)))))))))).)))))......---------------- ( -20.30, z-score = -2.15, R) >droWil1.scaffold_181017 627559 87 - 939937 UUGACAUUUCUUUAUGAGUUC-----UUAAACCACUUAACAAAAAUUUAAAGACCUAUAUUUAUAUACAAACUGAAAUUUGCUAUGUGUGUG-------- ...((((.((((((....((.-----((((.....)))).)).....)))))).........(((((((((......)))).))))))))).-------- ( -6.20, z-score = 1.02, R) >consensus UUUCCAUUUCCAUUUGGCUGU____CAUAAAUUACUUGCCAAAUUCCUUGGCAUUAACUUUGGCCGACAGCCCGGAAACAUAUAUGUG____________ ......(((((....((((((......((((.....((((((.....)))))).....))))....)))))).)))))...................... ( -6.96 = -10.03 + 3.06)


| Location | 11,242,062 – 11,242,162 |
|---|---|
| Length | 100 |
| Sequences | 4 |
| Columns | 100 |
| Reading direction | reverse |
| Mean pairwise identity | 63.57 |
| Shannon entropy | 0.55743 |
| G+C content | 0.37051 |
| Mean single sequence MFE | -23.17 |
| Consensus MFE | -12.30 |
| Energy contribution | -13.54 |
| Covariance contribution | 1.25 |
| Combinations/Pair | 1.32 |
| Mean z-score | -2.10 |
| Structure conservation index | 0.53 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.41 |
| SVM RNA-class probability | 0.990345 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 11242062 100 - 24543557 UGUAAGUAUGUACACAGUUAUGUUUCCGGGCUGUCGGCCAAAGUUAAUGCCAAGGAAUUUGGCAAGUAAUUUAUGACAGACAGCCAAAUGGAAAUGGAAA ((((......)))).......((((((.(((((((.(.((((((((.(((((((...)))))))..)))))).)).).)))))))....))))))..... ( -29.70, z-score = -2.70, R) >droSim1.chr3L 10636345 84 - 22553184 ------------CACAGAUAUGUUUCCGGGCUGUCGGCCAAAGUUAAUGCCAAGGAAUUUGGCAAGUAAUUUAUG----ACAGCCAAAUGGAAAUGGAAA ------------.........((((((.((((((((....((((((.(((((((...)))))))..)))))).))----))))))....))))))..... ( -28.60, z-score = -3.68, R) >droEre2.scaffold_4784 11252928 80 - 25762168 ----------------UAUAUGUUUCCGGGCUGUCGGCCAAAGUUAAUGCCAAGGAAUUUGCCAAGUAAUUUAUG----ACAGCCAAAUGGAAAUGGGAA ----------------.....((((((.((((((((((..........)))..((......))...........)----))))))....))))))..... ( -23.80, z-score = -1.96, R) >droWil1.scaffold_181017 627559 87 + 939937 --------CACACACAUAGCAAAUUUCAGUUUGUAUAUAAAUAUAGGUCUUUAAAUUUUUGUUAAGUGGUUUAA-----GAACUCAUAAAGAAAUGUCAA --------.....((((.((((((....)))))).......(((..((..((((((..((....))..))))))-----..))..))).....))))... ( -10.60, z-score = -0.04, R) >consensus ____________CACAGAUAUGUUUCCGGGCUGUCGGCCAAAGUUAAUGCCAAGGAAUUUGGCAAGUAAUUUAUG____ACAGCCAAAUGGAAAUGGAAA .....................((((((.((((((......((((((.(((((((...)))))))..)))))).......))))))....))))))..... (-12.30 = -13.54 + 1.25)


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:18:05 2011