
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 11,023,725 – 11,023,821 |
| Length | 96 |
| Max. P | 0.575076 |

| Location | 11,023,725 – 11,023,821 |
|---|---|
| Length | 96 |
| Sequences | 9 |
| Columns | 107 |
| Reading direction | forward |
| Mean pairwise identity | 79.18 |
| Shannon entropy | 0.41844 |
| G+C content | 0.42258 |
| Mean single sequence MFE | -24.36 |
| Consensus MFE | -16.88 |
| Energy contribution | -16.72 |
| Covariance contribution | -0.16 |
| Combinations/Pair | 1.26 |
| Mean z-score | -1.05 |
| Structure conservation index | 0.69 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.17 |
| SVM RNA-class probability | 0.575076 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 11023725 96 + 24543557 AAAUAAUAGCUAGGAGUAUG---------GAAGAGAGAUCUUCGAGGCAGCCAAUUUGUGUUCCGUUGCUUUGCCGUUUGUAUAAAUCAAUUGGCAAUGAGGCAA-- ........(((.........---------(((((....)))))((((((((.(((....)))..))))))))((((.(((.......))).)))).....)))..-- ( -21.90, z-score = -0.10, R) >droSim1.chr3L 10421194 96 + 22553184 AAAUAAUAGCUAGGAGUAUA---------GAAGAGAGAUCUUCGAGGCAGCCAAUUUGUGUUCCGUUGCUUUGCCGUUUGUAUAAAUCAAUUGGCAAUGAGGCAA-- ............((((((((---------(((((....)))))..((...))....)))))))).(((((((((((.(((.......))).))))))...)))))-- ( -22.90, z-score = -0.75, R) >droSec1.super_0 3251661 96 + 21120651 AAAUAAUAGCUAGGAGUAUA---------GAAGAGAGAUCUUCGAGGCAGCCAAUUUGUGUUCCGUUGCUUUGCCGUUUGUAUAAAUCAAUUGGCAAUGAGGCAA-- ............((((((((---------(((((....)))))..((...))....)))))))).(((((((((((.(((.......))).))))))...)))))-- ( -22.90, z-score = -0.75, R) >droYak2.chr3L 11051342 105 + 24197627 AAAUAAUAGCUAGAAGCAUAUAGCGCAGAGAAGAGAGAUCUUCGAGGCAGCCAAUUUGUGUUCCGUUGCUUUGCCGUUUGUAUAAAUCAAUUGGCAAUGAGGCAA-- ........(((.((((((...(((((((((((((....)))))..((...))..))))))))....))))))((((.(((.......))).)))).....)))..-- ( -28.60, z-score = -1.51, R) >droEre2.scaffold_4784 11033013 96 + 25762168 AAAUAAUAGCUUGGAGUAUA---------GAAGAGAGAUCUUCGAGGCAGCCAAUUUGUGUUCCGUUGCUUUGCCGUUUGUAUAAAUCAAUUGGCAAUGAGGCAA-- .....((((.((((.((...---------(((((....)))))...))..)))).))))......(((((((((((.(((.......))).))))))...)))))-- ( -23.30, z-score = -0.75, R) >droAna3.scaffold_13337 15937121 75 + 23293914 ------------------------------CGGUUUGGUCCGCGAGGCAACCAAUUUGUGUUCCGUUGCUUUGCCGUUUGUAUAAAUCAAUUGGCAAUGAGGCAA-- ------------------------------(((.(((((((....))..)))))........)))(((((((((((.(((.......))).))))))...)))))-- ( -21.20, z-score = -0.80, R) >dp4.chrXR_group6 7309271 98 - 13314419 AGAGUGUAGAGUGGAGAGGG---------AGAGAGAUCUCUUCGAGGCAACCAAUUUGUGUUCCGUUGCUUUGCCGUUUGUAUAAAUCAAUUGGCAAUGAGACGAUG .(((..((((.(((....((---------((((....)))))).......))).))))..)))((((...((((((.(((.......))).))))))...))))... ( -28.20, z-score = -1.80, R) >droPer1.super_47 89979 98 - 592741 AGAGUGUAGAGUGGAGAGGG---------AGAGAGAUCUCUUCGAGGCAACCAAUUUGUGUUCCGUUGCUUUGCCGUUUGUAUAAAUCAAUUGGCAAUGAGACGAUG .(((..((((.(((....((---------((((....)))))).......))).))))..)))((((...((((((.(((.......))).))))))...))))... ( -28.20, z-score = -1.80, R) >droWil1.scaffold_180916 2557279 85 + 2700594 -------AGCCAACAGUC-----------ACUGAGAUCU-UUUGAGGCAACCAAUUUGUGUUCCGUUGCCUUGCCGUUUGUAUAAAUCAAUUGGCUAGUGGACU--- -------.......((((-----------((((......-...((((((((.(((....)))..))))))))((((.(((.......))).)))))))).))))--- ( -22.00, z-score = -1.18, R) >consensus AAAUAAUAGCUAGGAGUAUA_________GAAGAGAGAUCUUCGAGGCAGCCAAUUUGUGUUCCGUUGCUUUGCCGUUUGUAUAAAUCAAUUGGCAAUGAGGCAA__ .............................(((((....)))))((((((((.(((....)))..))))))))((((.(((.......))).))))............ (-16.88 = -16.72 + -0.16)

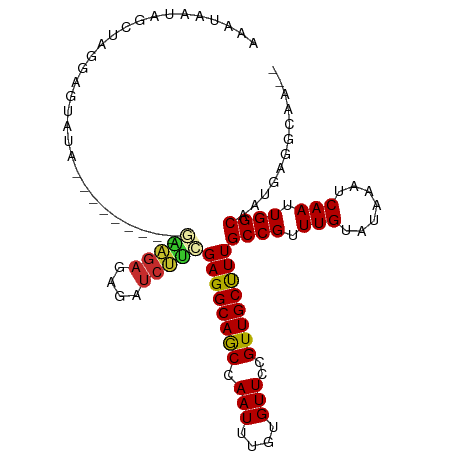

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:17:35 2011