
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 10,982,752 – 10,982,860 |
| Length | 108 |
| Max. P | 0.579737 |

| Location | 10,982,752 – 10,982,860 |
|---|---|
| Length | 108 |
| Sequences | 8 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 82.83 |
| Shannon entropy | 0.34484 |
| G+C content | 0.39386 |
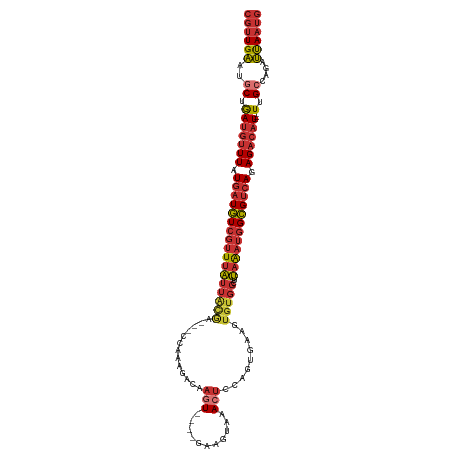
| Mean single sequence MFE | -27.04 |
| Consensus MFE | -13.76 |
| Energy contribution | -14.54 |
| Covariance contribution | 0.78 |
| Combinations/Pair | 1.22 |
| Mean z-score | -2.09 |
| Structure conservation index | 0.51 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.18 |
| SVM RNA-class probability | 0.579737 |
| Prediction | RNA |
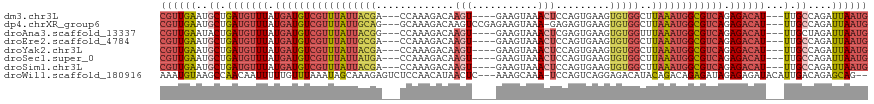
Download alignment: ClustalW | MAF
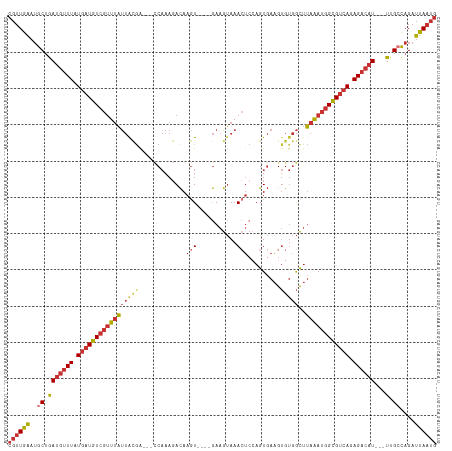
>dm3.chr3L 10982752 108 - 24543557 CGUUGAAUGCUGAUGUUUAUGAUGUCGUUUAUUACGA---CCAAAGACAAGU----GAAGUAAACUCCAGUGAAGUGUGGCUUAAAUGGCGUCAGAGACAU---UUGCCAGAUUAAUG ((((((..((.(((((((.((((((((((((....(.---(((......(.(----(.((....)).)).)......)))).)))))))))))).))))))---).))....)))))) ( -28.30, z-score = -2.16, R) >dp4.chrXR_group6 7264673 111 + 13314419 CGUUGAAUGCUGAUGUUUAUGAUGUCGUUUAUUGCAG---GCAAAGACAAGUCCGAGAAGUAAA-GAGAGUGAAGUGUGGCUUAAAUGGCGUCAGAGACAU---UUGCCAGAUUAAUG ((((((..((.(((((((.((((((((((((..((..---(((......(.(((..........-).)).)....))).)).)))))))))))).))))))---).))....)))))) ( -28.40, z-score = -1.86, R) >droAna3.scaffold_13337 15895471 108 - 23293914 CGUUGAAUACUGAUGUUUAUGAUGUCGUUUAUUACGG---CCAAAGACAAGU----GAAGUAAACUCGAGUGAAGUGUGGUUUAAAUGGCGUCAGAGACAU---UUGCUAGAUUAAUG ((((((.(((.(((((((.((((((((((((....((---(((....((..(----((.......)))..)).....))))))))))))))))).))))))---).).))..)))))) ( -27.70, z-score = -2.10, R) >droEre2.scaffold_4784 10990786 108 - 25762168 CGUUGAAUGCUGAUGUUUAUGAUGUCGUUUAUUGCGA---CCAAAGACAAGU----GAAGUAAACUCCAGUGAAGUGUGGCUUAAAUGGCGUCAGAGACAU---UUGCCAGAUUAAUG ((((((..((.(((((((.((((((((((((..((.(---(........(.(----(.((....)).)).).....)).)).)))))))))))).))))))---).))....)))))) ( -29.22, z-score = -2.25, R) >droYak2.chr3L 11008925 108 - 24197627 CGUUGAAUGCUGAUGUUUAUGAUGUCGUUUAUUACGA---CCAAAGACAAGU----GAAGUAAACUCCAGUGAAGUGUGGCUUAAAUGGCGUCAGAGACAU---UUGCCAGAUUAAUG ((((((..((.(((((((.((((((((((((....(.---(((......(.(----(.((....)).)).)......)))).)))))))))))).))))))---).))....)))))) ( -28.30, z-score = -2.16, R) >droSec1.super_0 3206285 108 - 21120651 CGUUGAAUGCUGAUGUUUAUGAUGUCGUUUAUUAUGA---CCAAAGACAAGU----GAAGUAAACUCCAGUGAAGUGUGGCUUAAAUGGCGUCAGAGACAU---UUGCCAGAUUAAUG ((((((..((.(((((((.((((((((((((....(.---(((......(.(----(.((....)).)).)......)))).)))))))))))).))))))---).))....)))))) ( -29.00, z-score = -2.58, R) >droSim1.chr3L 10378171 108 - 22553184 CGUUGAAUGCUGAUGUUUAUGAUGUCGUUUAUUACGA---CCAAAGACAAGU----GAAGUAAACUCCAGUGAAGUGUGGCUUAAAUGGCGUCAGAGACAU---UUGCCAGAUUAAUG ((((((..((.(((((((.((((((((((((....(.---(((......(.(----(.((....)).)).)......)))).)))))))))))).))))))---).))....)))))) ( -28.30, z-score = -2.16, R) >droWil1.scaffold_180916 79896 112 + 2700594 AAAUGUAAGCCAACAAUUUUUGUUUAAAUAGCAAAGAGUCUCCAACAUAACUC---AAAGCAAA-UCCAGUCAGGAGACAUACAGACAGAGAUAGAGAGAUACAUUGACAGAGCAG-- .((((((.........((((((((.....))))))))((((((..........---........-........)))))).....................))))))..........-- ( -17.07, z-score = -1.46, R) >consensus CGUUGAAUGCUGAUGUUUAUGAUGUCGUUUAUUACGA___CCAAAGACAAGU____GAAGUAAACUCCAGUGAAGUGUGGCUUAAAUGGCGUCAGAGACAU___UUGCCAGAUUAAUG ((((((...(((((((((.((((((((((((...................................................)))))))))))).)))))).......))).)))))) (-13.76 = -14.54 + 0.78)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:17:31 2011