
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 10,835,227 – 10,835,339 |
| Length | 112 |
| Max. P | 0.500000 |

| Location | 10,835,227 – 10,835,339 |
|---|---|
| Length | 112 |
| Sequences | 8 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 69.29 |
| Shannon entropy | 0.61136 |
| G+C content | 0.51858 |
| Mean single sequence MFE | -24.45 |
| Consensus MFE | -11.15 |
| Energy contribution | -11.34 |
| Covariance contribution | 0.19 |
| Combinations/Pair | 1.42 |
| Mean z-score | -0.92 |
| Structure conservation index | 0.46 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.02 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
>dm3.chr3L 10835227 112 + 24543557 ------CAUCCCGUUCCGAAUCCCUCCCUCAGAGUCACCCACUCCAGUCCACGGAACAACUGCGUAUGACUAAUCUUUGACGGUGGCUUUUGUUGUCGUCAUCG--UCUUUGCAAACGAA ------......((((((.....(((.....)))......((....))...))))))...((((...(((.......((((((..((....))..))))))..)--))..))))...... ( -26.60, z-score = -1.06, R) >droSim1.chr3L 10229391 112 + 22553184 ------CAUCCCGUUCCGAAUCCCUCCCUCAGAGUCACCCACUCCAGUCCACGGAACAACUGCGUAUGACUAAUCUUUGACGGUGGCUUUUGUUGUCGUCAUCG--UCUUUGCAAACGAA ------......((((((.....(((.....)))......((....))...))))))...((((...(((.......((((((..((....))..))))))..)--))..))))...... ( -26.60, z-score = -1.06, R) >droSec1.super_0 3055669 112 + 21120651 ------CAUCCCGUUCCGAAUCCCUCCCUCAGAGUCACCCACUCCAGUCCACGGAACAACUGCGUAUGACUAAUCUUUGACGGUGGCUUUUGUUGUCGUCAUCG--UCUUUGCAAACGAA ------......((((((.....(((.....)))......((....))...))))))...((((...(((.......((((((..((....))..))))))..)--))..))))...... ( -26.60, z-score = -1.06, R) >droYak2.chr3L 10852608 118 + 24197627 CCAUCUCCUUCCGUUCCGAAUCCCUCCCUCAGAGUCACCCACUCCAGUCCGCGGAACAACUGCGUAUGACUAAUCUUUGACGGUGGCUUUUGUUGUCGUCAUCG--UCUUUGCCAACGAA ...........((((..((.....))...(((((.(.........(((((((((.....)))))...))))......((((((..((....))..))))))..)--.)))))..)))).. ( -26.40, z-score = -0.78, R) >droEre2.scaffold_4784 10838434 112 + 25762168 ------CUUUCCGUUCCGAAUCCCUCCCUCAGAGUCACCCACUCCAGUCCACGGAACAACUGCGUAUGACUAAUCCUUGACGGUGGCUUUUGUUGUCGUCAUCG--UCUUUGCCAGCGAA ------......((((((.....(((.....)))......((....))...))))))..(((.(((.(((.......((((((..((....))..))))))..)--))..)))))).... ( -27.00, z-score = -0.98, R) >droAna3.scaffold_13337 15746833 84 + 23293914 -------------------------ACCCCCGCUUCCUACGGGCCAGCAUUCGGGCCAACUGUGUAUGACUAAUCUUUGACGGUGGCUUUUGUUGUCGUCAUCG--UCUUU--------- -------------------------.(((..(((.((....))..)))....)))............(((.......((((((..((....))..))))))..)--))...--------- ( -21.90, z-score = -0.46, R) >dp4.chrXR_group6 7096491 103 - 13314419 ------------ACUCCCACUCCCUCCCCCUCCCUCCUCC-CUUGGGCACUCGCUG----UGUGUAUGACUAAUCUUUGACGGUGGCUUUUGUUGUCGUCAUCGCGUCUUGGCGAACCAA ------------............................-..(((((((.....)----)))..............((((((..((....))..))))))((((......)))).))). ( -23.80, z-score = -1.10, R) >droGri2.scaffold_15110 4077583 87 - 24565398 ---------------------CUAACCACCCCCACUUAGAGCGCCAGGCAACAACAAUACUUUGUAUGACUAAUCUUUGGCG-------UCGUCGUCGCCAUCAUGACAAGACAA----- ---------------------.................((.((((((....).((((....))))............)))))-------))(((((((......))))..)))..----- ( -16.70, z-score = -0.85, R) >consensus ______C_U_CCGUUCCGAAUCCCUCCCUCAGAGUCACCCACUCCAGUCCACGGAACAACUGCGUAUGACUAAUCUUUGACGGUGGCUUUUGUUGUCGUCAUCG__UCUUUGCAAACGAA ...............................((((.....)))).................................((((((..((....))..))))))................... (-11.15 = -11.34 + 0.19)
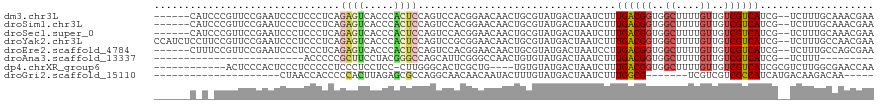


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:17:07 2011