| Sequence ID | dm3.chr3L |
|---|---|
| Location | 10,829,420 – 10,829,483 |
| Length | 63 |
| Max. P | 0.923887 |

| Location | 10,829,420 – 10,829,483 |
|---|---|
| Length | 63 |
| Sequences | 11 |
| Columns | 63 |
| Reading direction | forward |
| Mean pairwise identity | 93.19 |
| Shannon entropy | 0.15867 |
| G+C content | 0.32586 |
| Mean single sequence MFE | -13.40 |
| Consensus MFE | -12.06 |
| Energy contribution | -12.07 |
| Covariance contribution | 0.01 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.89 |
| Structure conservation index | 0.90 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.30 |
| SVM RNA-class probability | 0.923887 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 10829420 63 + 24543557 GGCUAUUUGCAUACUUAAGUGUGCAGUUAUUACCUUGUUAUAUCGAUAUAAAUAACAACUGCG .((....(((((((....)))))))((((((...(((......)))....))))))....)). ( -13.50, z-score = -1.75, R) >droSim1.chr3L 10223414 63 + 22553184 GGCUAUUUGCAUACUUAAGUGUGCAGUUAUUACCUUGUUAUAUCGAUAUAAAUAACAACUGCG .((....(((((((....)))))))((((((...(((......)))....))))))....)). ( -13.50, z-score = -1.75, R) >droSec1.super_0 3049926 63 + 21120651 GGCUAUUUGCAUACUUAAGUGUGCAGUUAUUACCUUGUUAUAUCGAUAUAAAUAACAACUGCG .((....(((((((....)))))))((((((...(((......)))....))))))....)). ( -13.50, z-score = -1.75, R) >droYak2.chr3L 10846813 63 + 24197627 GGCUAUUUGCAUACUUAAGUGUGCAGUUAUUACCUUGUUAUAUCGAUAUAAAUAACAACUGCG .((....(((((((....)))))))((((((...(((......)))....))))))....)). ( -13.50, z-score = -1.75, R) >droEre2.scaffold_4784 10832863 63 + 25762168 GGCUAUUUGCAUACUUAAGUGUGCAGUUAUUACCUUGUUAUAUCGAUAUAAAUAACAACUGCG .((....(((((((....)))))))((((((...(((......)))....))))))....)). ( -13.50, z-score = -1.75, R) >droAna3.scaffold_13337 15741879 63 + 23293914 GGCUAUUUGCAUACUUAAGUGUGCAGUUAUUACCCUGUUAUAUCGAUAUAAAUAACAACAAUU ((.((.((((((((....))))))))....)).))((((((..........))))))...... ( -12.30, z-score = -1.80, R) >dp4.chrXR_group6 7089665 63 - 13314419 GGCUAUUUGCAUACUUAAGUGUGCAGUUAUUACCUUGUUAUAUCGAUAUAAAUAACAACUGAG ((....((((((((....))))))))......))(((((((..........)))))))..... ( -12.50, z-score = -1.55, R) >droPer1.super_36 16095 63 + 818889 GGCUAUUUGCAUACUUAAGUGUGCAGUUAUUACCUUGUUAUAUCGAUAUAAAUAACAACUGAG ((....((((((((....))))))))......))(((((((..........)))))))..... ( -12.50, z-score = -1.55, R) >droWil1.scaffold_180949 250292 58 - 6375548 GGCCAUUUGCAUAUUUAAGUGUGCAGUUAUUACCUUGUUAUUUCGAUAUAAAUAACAA----- ((....((((((((....))))))))......)).((((((((......)))))))).----- ( -12.50, z-score = -2.13, R) >droVir3.scaffold_13049 15310888 63 - 25233164 GGCUAUUUGCAUACUUAAGUGUGCAGUUAUUACCUUGUUAUUCCGAUAUAAAUAACAAUGACG .......(((((((....)))))))(((((.....(((((((........)))))))))))). ( -14.40, z-score = -2.63, R) >droMoj3.scaffold_6680 10856526 62 + 24764193 GGCUAUUUGCAUACUUAAGUGUGCAGUUAUUACCUUGUUAUUCCGUCAUAAAUAACAAUGGC- ......((((((((....))))))))......((((((((((........)))))))).)).- ( -15.70, z-score = -2.37, R) >consensus GGCUAUUUGCAUACUUAAGUGUGCAGUUAUUACCUUGUUAUAUCGAUAUAAAUAACAACUGCG ((....((((((((....))))))))......))(((((((..........)))))))..... (-12.06 = -12.07 + 0.01)

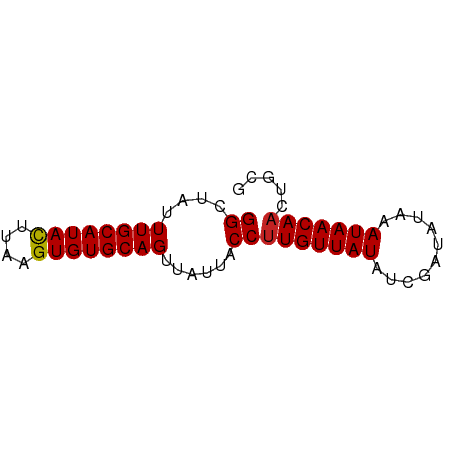
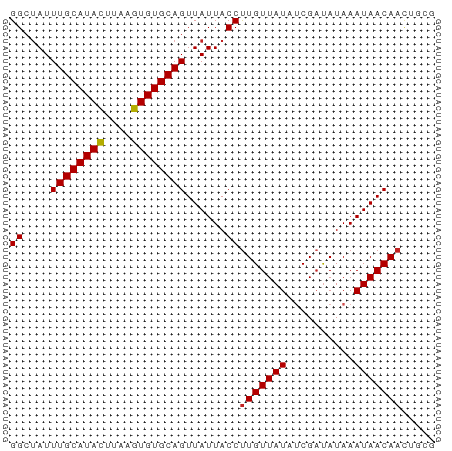
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:17:06 2011