| Sequence ID | dm3.chr3L |
|---|---|
| Location | 10,744,292 – 10,744,388 |
| Length | 96 |
| Max. P | 0.904152 |

| Location | 10,744,292 – 10,744,388 |
|---|---|
| Length | 96 |
| Sequences | 9 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 90.31 |
| Shannon entropy | 0.20050 |
| G+C content | 0.35269 |
| Mean single sequence MFE | -21.39 |
| Consensus MFE | -15.77 |
| Energy contribution | -16.88 |
| Covariance contribution | 1.11 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.40 |
| Structure conservation index | 0.74 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.17 |
| SVM RNA-class probability | 0.904152 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
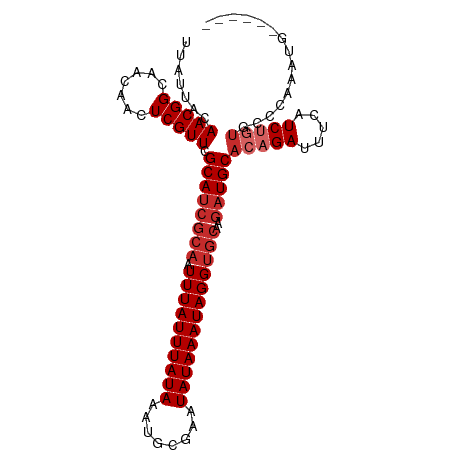
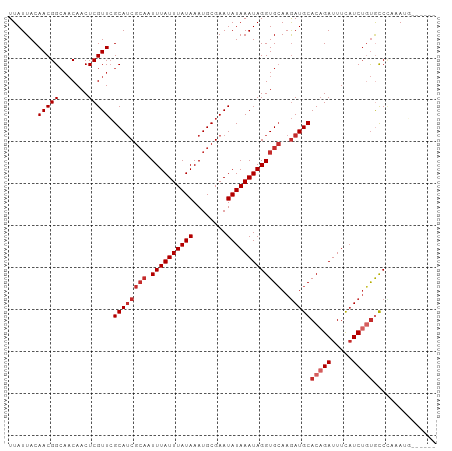
>dm3.chr3L 10744292 96 - 24543557 UUAUUACAACGGCAACAACUCGUUCGCAUCGCAAUUUAUUUAUAAAUGCGAAUAUAAAUAGGUGCAAGAUGCACAGAUUUCAUCUGUGCCCAAAUG------ ..........(((............((((((((.((((((((((........)))))))))))))..)))))((((((...)))))))))......------ ( -24.10, z-score = -2.95, R) >droSim1.chr3L 10137641 96 - 22553184 UUAUUACAACGGCAACAACUCGUUCGCAUCGCAAUUUAUUUAUAAAUGCGAAUAUAAAUAGGUGCAAGAUGCACAGAUUUCAUCUGUGCCCAAAUG------ ..........(((............((((((((.((((((((((........)))))))))))))..)))))((((((...)))))))))......------ ( -24.10, z-score = -2.95, R) >droSec1.super_0 2964521 96 - 21120651 UUAUUACAACGGCAACAACUCGUUCGCAUCGCAAUUUAUUUAUAAAUGCGAAUAUAAAUAGGUGCAAGAUGCACUGAUUUCAUCUGUGCCCAAAUG------ ..........((((.((....((((((((................))))))))..((((.(((((.....))))).))))....))))))......------ ( -20.59, z-score = -1.53, R) >droYak2.chr3L 10756671 96 - 24197627 UUAUUACAACGGCAACAACUCGUUCGCAUCGCAAUUUAUUUAUAAAUGCGAAUAUAAAUAGGUGCAAGAUGCACAGAUUUCAUCUGUGCCCAAAAG------ ..........(((............((((((((.((((((((((........)))))))))))))..)))))((((((...)))))))))......------ ( -24.10, z-score = -3.27, R) >droEre2.scaffold_4784 10746892 96 - 25762168 UUAUUACAACGGCAACAACUCGUUCGCAACGCAAUUUAUUUAUAAAUGCGAAUAUAAAUAGGUGCAAGAUGCACAGAUUUCAUCUGUGCCCAAAUG------ ...........(((((.....))).))...(((.((((((((((........))))))))))))).....((((((((...)))))))).......------ ( -21.20, z-score = -1.97, R) >droAna3.scaffold_13337 15657249 93 - 23293914 UUAUUACAACGGCAACAACUCGUUCGCAUCGCAUUUUAUUUAUAAAUGCGAAUAUAAAUAGGUGCAAGAUGCACAGAUUUCAUC--UGCCUAAAA------- ..........((((..............((((((((.......))))))))....((((..((((.....))))..))))....--)))).....------- ( -21.40, z-score = -2.74, R) >dp4.chrXR_group6 6999854 94 + 13314419 UUAUUACAACGGCAACAACUCGUUCGCAUCGCAUUUUAUUUAUAAAUGCGAAUAUAAAUAGGUGCAAGAUGCACAGAUUUCAUCUGUAUCAUCA-------- ..........(....).........(((((((((((.(((((((........)))))))))))))..)))))((((((...)))))).......-------- ( -20.50, z-score = -2.60, R) >droPer1.super_940 5545 94 - 11099 UUAUUACAACGGCAACAACUCGUUCGCAUCGCAUUUUAUUUAUAAAUGCGAAUAUAAAUAGGUGCAAGAUGCACAGAUUUCAUCUGUAUCAUCA-------- ..........(....).........(((((((((((.(((((((........)))))))))))))..)))))((((((...)))))).......-------- ( -20.50, z-score = -2.60, R) >droWil1.scaffold_180949 109284 95 + 6375548 UUAUUACAACGG---CAACUCGUUGGCAUCG---UUUAUUUAUAAAUGCGAAUAUAAAUAGGUGGAAAAUGCACAGAUUUU-UCUUCGUCUUUAUCUCAUUG ......((((((---....))))))......---.......(((((.(((((...((((..(((.......)))..)))).-..))))).)))))....... ( -16.00, z-score = -0.97, R) >consensus UUAUUACAACGGCAACAACUCGUUCGCAUCGCAAUUUAUUUAUAAAUGCGAAUAUAAAUAGGUGCAAGAUGCACAGAUUUCAUCUGUGCCCAAAUG______ .......(((((.......))))).((((((((.((((((((((........)))))))))))))..)))))(((((.....)))))............... (-15.77 = -16.88 + 1.11)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:16:57 2011