
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 10,701,736 – 10,701,828 |
| Length | 92 |
| Max. P | 0.870473 |

| Location | 10,701,736 – 10,701,828 |
|---|---|
| Length | 92 |
| Sequences | 8 |
| Columns | 96 |
| Reading direction | forward |
| Mean pairwise identity | 77.46 |
| Shannon entropy | 0.41511 |
| G+C content | 0.49185 |
| Mean single sequence MFE | -21.75 |
| Consensus MFE | -10.79 |
| Energy contribution | -12.17 |
| Covariance contribution | 1.38 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.08 |
| Structure conservation index | 0.50 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.79 |
| SVM RNA-class probability | 0.818722 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
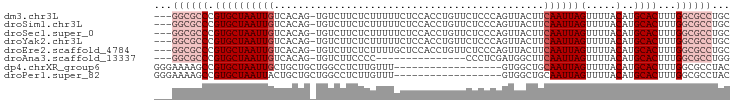
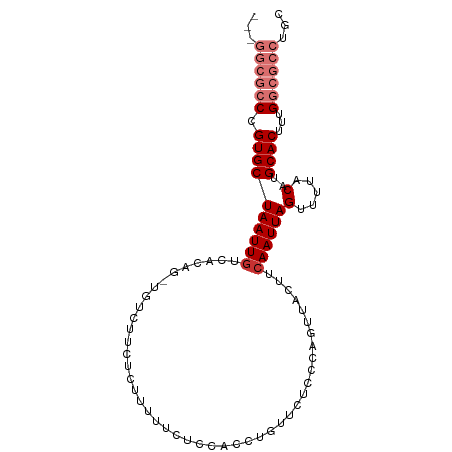
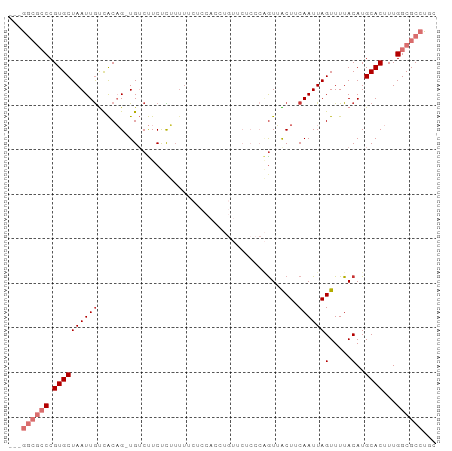
>dm3.chr3L 10701736 92 + 24543557 ---GGCGCCCGUGCUAAUUGUCACAG-UGUCUUCUCUUUUUCUCCACCUGUUCUCCCAGUUACUUCAAUUAGUUUUACAUGCACUUUGGCGCCUGC ---((((((.((((........((((-.((...............)))))).......((.(((......)))...))..))))...))))))... ( -20.86, z-score = -2.81, R) >droSim1.chr3L 10094622 92 + 22553184 ---GGCGCCCGUGCUAAUUGUCACAG-UGUCUUCUCUUUUUCUCCACCUGUUCUCCCAGUUACUUCAAUUAGUUUUACAUGCACUUUGGCGCCUGC ---((((((.((((........((((-.((...............)))))).......((.(((......)))...))..))))...))))))... ( -20.86, z-score = -2.81, R) >droSec1.super_0 2922444 92 + 21120651 ---GGCGCCCGUGCUAAUUGUCACAG-UGUCUUCUCUUUUUCUCCACCUGUUCUCCCAGUUACUUCAAUUAGUUUUACAUGCACUUUGGCGCCUGC ---((((((.((((........((((-.((...............)))))).......((.(((......)))...))..))))...))))))... ( -20.86, z-score = -2.81, R) >droYak2.chr3L 10712052 92 + 24197627 ---GGCGCCCGUGCUAAUUGUCACAG-UGUCUUCUCUUUUUCUCCACCUGUUCUCCCAGUUACUUCAAUUAGUUUUACAUGCACUUUGGCGCCUGC ---((((((.((((........((((-.((...............)))))).......((.(((......)))...))..))))...))))))... ( -20.86, z-score = -2.81, R) >droEre2.scaffold_4784 10700630 92 + 25762168 ---GGCGCCCGUGCUAAUUGUCACAG-UGUCUUCUCUUUUGCUCCACCUGUUCUCCCAGUUACUUCAAUUAGUUUUACAUGCACUUUGGCGCCUGC ---((((((.((((........((((-.((...(......)....)))))).......((.(((......)))...))..))))...))))))... ( -20.90, z-score = -1.99, R) >droAna3.scaffold_13337 15613525 77 + 23293914 ---GGCGCCCGUGCUAAUUGUCACAG-UGUCUUCCCC---------------CCCUCGAUGGCUUCAAUUAGUUUUACAUGCACUUUGGCGCCUGG ---((((((.((((((((((......-.(((.((...---------------.....)).)))..))))))(.....)..))))...))))))... ( -20.60, z-score = -1.23, R) >dp4.chrXR_group6 6982737 78 - 13314419 GGGAAAAGCCGUGCUAAUUGCUGCUGCUGGCCUCUUGUUU------------------GUGGCUGCAAUUAGUUUUACAUGCACUUUGGCGCCUAC (((....((((.(((((((((.((..(..((.....))..------------------)..)).))))))))).............)))).))).. ( -25.94, z-score = -1.34, R) >droPer1.super_82 267072 78 + 267963 GGGAAAAGCCGUGCUAAUUACUGCUGCUGGCCUCUUGUUU------------------GUGGCUGCAAUUAGUUUUACAUGCACUUUGGCGCCUAC (((....(((((((.....((((.(((.((((.(......------------------).)))))))..)))).......))))...))).))).. ( -23.10, z-score = -0.83, R) >consensus ___GGCGCCCGUGCUAAUUGUCACAG_UGUCUUCUCUUUUUCUCCACCUGUUCUCCCAGUUACUUCAAUUAGUUUUACAUGCACUUUGGCGCCUGC ...((((((.((((((((((.............................................))))))(.....)..))))...))))))... (-10.79 = -12.17 + 1.38)
| Location | 10,701,736 – 10,701,828 |
|---|---|
| Length | 92 |
| Sequences | 8 |
| Columns | 96 |
| Reading direction | reverse |
| Mean pairwise identity | 77.46 |
| Shannon entropy | 0.41511 |
| G+C content | 0.49185 |
| Mean single sequence MFE | -23.04 |
| Consensus MFE | -14.03 |
| Energy contribution | -13.86 |
| Covariance contribution | -0.17 |
| Combinations/Pair | 1.24 |
| Mean z-score | -1.76 |
| Structure conservation index | 0.61 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.00 |
| SVM RNA-class probability | 0.870473 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
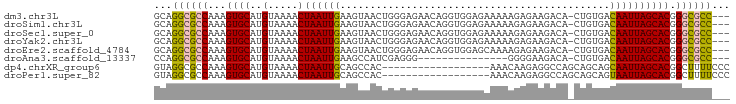
>dm3.chr3L 10701736 92 - 24543557 GCAGGCGCCAAAGUGCAUGUAAAACUAAUUGAAGUAACUGGGAGAACAGGUGGAGAAAAAGAGAAGACA-CUGUGACAAUUAGCACGGGCGCC--- ...((((((...((((..(.....)((((((......(((......)))(((...............))-).....)))))))))).))))))--- ( -25.66, z-score = -2.95, R) >droSim1.chr3L 10094622 92 - 22553184 GCAGGCGCCAAAGUGCAUGUAAAACUAAUUGAAGUAACUGGGAGAACAGGUGGAGAAAAAGAGAAGACA-CUGUGACAAUUAGCACGGGCGCC--- ...((((((...((((..(.....)((((((......(((......)))(((...............))-).....)))))))))).))))))--- ( -25.66, z-score = -2.95, R) >droSec1.super_0 2922444 92 - 21120651 GCAGGCGCCAAAGUGCAUGUAAAACUAAUUGAAGUAACUGGGAGAACAGGUGGAGAAAAAGAGAAGACA-CUGUGACAAUUAGCACGGGCGCC--- ...((((((...((((..(.....)((((((......(((......)))(((...............))-).....)))))))))).))))))--- ( -25.66, z-score = -2.95, R) >droYak2.chr3L 10712052 92 - 24197627 GCAGGCGCCAAAGUGCAUGUAAAACUAAUUGAAGUAACUGGGAGAACAGGUGGAGAAAAAGAGAAGACA-CUGUGACAAUUAGCACGGGCGCC--- ...((((((...((((..(.....)((((((......(((......)))(((...............))-).....)))))))))).))))))--- ( -25.66, z-score = -2.95, R) >droEre2.scaffold_4784 10700630 92 - 25762168 GCAGGCGCCAAAGUGCAUGUAAAACUAAUUGAAGUAACUGGGAGAACAGGUGGAGCAAAAGAGAAGACA-CUGUGACAAUUAGCACGGGCGCC--- ...((((((...((((..(.....)((((((......(((......)))(((...(......)....))-).....)))))))))).))))))--- ( -25.80, z-score = -2.33, R) >droAna3.scaffold_13337 15613525 77 - 23293914 CCAGGCGCCAAAGUGCAUGUAAAACUAAUUGAAGCCAUCGAGGG---------------GGGGAAGACA-CUGUGACAAUUAGCACGGGCGCC--- ...((((((...((((........((..((((.....))))..)---------------)((.......-))..........)))).))))))--- ( -22.90, z-score = -1.20, R) >dp4.chrXR_group6 6982737 78 + 13314419 GUAGGCGCCAAAGUGCAUGUAAAACUAAUUGCAGCCAC------------------AAACAAGAGGCCAGCAGCAGCAAUUAGCACGGCUUUUCCC ((.(((((......)).(((((......))))))))))------------------.....(((((((....((........))..)))))))... ( -16.50, z-score = 0.74, R) >droPer1.super_82 267072 78 - 267963 GUAGGCGCCAAAGUGCAUGUAAAACUAAUUGCAGCCAC------------------AAACAAGAGGCCAGCAGCAGUAAUUAGCACGGCUUUUCCC ((.(((((......)).(((((......))))))))))------------------.....(((((((....((........))..)))))))... ( -16.50, z-score = 0.49, R) >consensus GCAGGCGCCAAAGUGCAUGUAAAACUAAUUGAAGUAACUGGGAGAACAGGUGGAGAAAAAGAGAAGACA_CUGUGACAAUUAGCACGGGCGCC___ ...((((((...((((..(.....)((((((.............................................)))))))))).))))))... (-14.03 = -13.86 + -0.17)

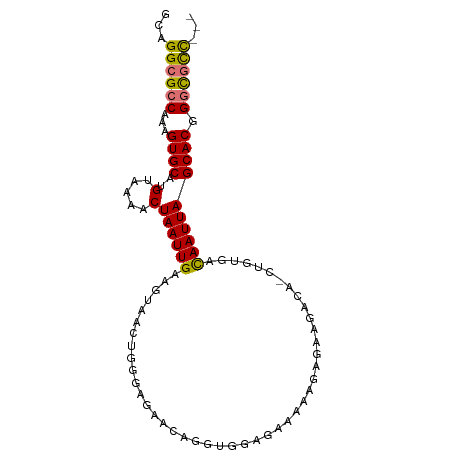
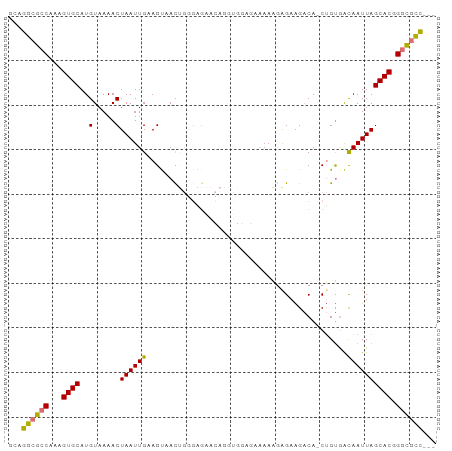
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:16:52 2011