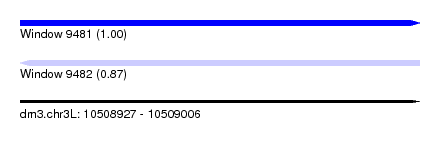

| Sequence ID | dm3.chr3L |
|---|---|
| Location | 10,508,927 – 10,509,006 |
| Length | 79 |
| Max. P | 0.996701 |

| Location | 10,508,927 – 10,509,006 |
|---|---|
| Length | 79 |
| Sequences | 5 |
| Columns | 87 |
| Reading direction | forward |
| Mean pairwise identity | 79.69 |
| Shannon entropy | 0.35512 |
| G+C content | 0.31744 |
| Mean single sequence MFE | -15.92 |
| Consensus MFE | -11.80 |
| Energy contribution | -13.12 |
| Covariance contribution | 1.32 |
| Combinations/Pair | 1.18 |
| Mean z-score | -2.92 |
| Structure conservation index | 0.74 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.97 |
| SVM RNA-class probability | 0.996701 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
>dm3.chr3L 10508927 79 + 24543557 --------UUCAUUUUCAAACUUUUAAAAUUGCUUAACAUUUCGAUAAAUUCUUAUCGAUAACAGAAUGUUAAGCAACAUAGCGCAU --------.....................(((((((((((((((((((....)))))))......)))))))))))).......... ( -18.20, z-score = -4.54, R) >droEre2.scaffold_4784 10503144 87 + 25762168 UAUCAAGAUUCGUGUUCAAAAUUUCGAAAAUGCCUAACAUUCCGAUACAUUGCUAUCGAUAACACAGUGUUAGGCAACAGAGCACAU ...........((((((....((....)).(((((((((((.(((((......))))).......)))))))))))...)))))).. ( -21.30, z-score = -3.35, R) >droYak2.chr3L 10502227 79 + 24197627 --------UUUAUUUUCAAAUUUUCGAAACUGCCUAACAUUCCGAUACAUUCCUAUCGAUAACACACUGUUAAGCAACAGAGCACAU --------.....((((........)))).(((.........(((((......)))))........(((((....))))).)))... ( -9.30, z-score = -2.14, R) >droSec1.super_0 2724491 85 + 21120651 --UUUUAAUCCAUUUUCAAACUUUUGAAAUAGCUUAACAUUUCGAUAAAUUCUUAUCGAUAACAGCGUGUUAAGCAACAGAGCGCAU --...........((((((....))))))..(((((((((.(((((((....))))))).......)))))))))............ ( -17.80, z-score = -3.12, R) >droSim1.chr3L 9905461 85 + 22553184 --UUUUAAUACAUUUUCAAACUUUCGAAAUAACUUAACAUUUCGAUAGAUGCUUAUCGAUAACAGCAUGUUAAGCAACAGAGCGCAU --..................((.(((((((........))))))).))((((.......((((.....)))).((......)))))) ( -13.00, z-score = -1.46, R) >consensus __U____AUUCAUUUUCAAACUUUCGAAAUUGCUUAACAUUUCGAUAAAUUCUUAUCGAUAACAGAAUGUUAAGCAACAGAGCGCAU ...............................(((((((((((((((((....)))))))......))))))))))............ (-11.80 = -13.12 + 1.32)
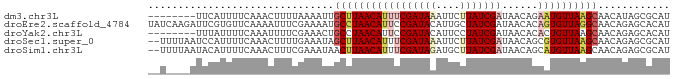
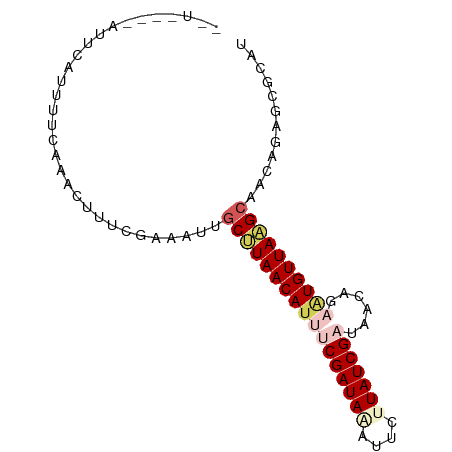
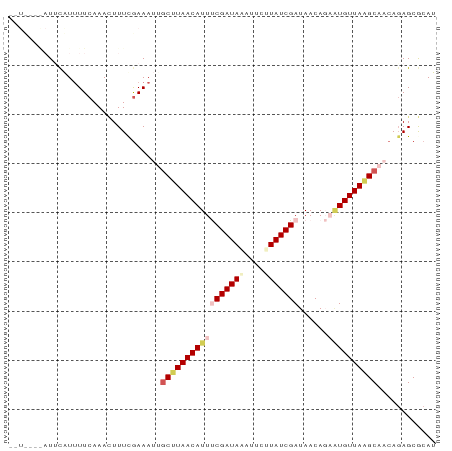
| Location | 10,508,927 – 10,509,006 |
|---|---|
| Length | 79 |
| Sequences | 5 |
| Columns | 87 |
| Reading direction | reverse |
| Mean pairwise identity | 79.69 |
| Shannon entropy | 0.35512 |
| G+C content | 0.31744 |
| Mean single sequence MFE | -17.06 |
| Consensus MFE | -14.28 |
| Energy contribution | -13.28 |
| Covariance contribution | -1.00 |
| Combinations/Pair | 1.30 |
| Mean z-score | -1.31 |
| Structure conservation index | 0.84 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.99 |
| SVM RNA-class probability | 0.869263 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
>dm3.chr3L 10508927 79 - 24543557 AUGCGCUAUGUUGCUUAACAUUCUGUUAUCGAUAAGAAUUUAUCGAAAUGUUAAGCAAUUUUAAAAGUUUGAAAAUGAA-------- .........(((((((((((((......(((((((....))))))))))))))))))))....................-------- ( -18.40, z-score = -2.57, R) >droEre2.scaffold_4784 10503144 87 - 25762168 AUGUGCUCUGUUGCCUAACACUGUGUUAUCGAUAGCAAUGUAUCGGAAUGUUAGGCAUUUUCGAAAUUUUGAACACGAAUCUUGAUA .((((......(((((((((.....((.((((((......)))))))))))))))))..(((((....))))))))).......... ( -18.10, z-score = -0.51, R) >droYak2.chr3L 10502227 79 - 24197627 AUGUGCUCUGUUGCUUAACAGUGUGUUAUCGAUAGGAAUGUAUCGGAAUGUUAGGCAGUUUCGAAAAUUUGAAAAUAAA-------- ..........((((((((((........((((((......))))))..))))))))))((((((....)))))).....-------- ( -15.40, z-score = -0.99, R) >droSec1.super_0 2724491 85 - 21120651 AUGCGCUCUGUUGCUUAACACGCUGUUAUCGAUAAGAAUUUAUCGAAAUGUUAAGCUAUUUCAAAAGUUUGAAAAUGGAUUAAAA-- ......((((((((((((((........(((((((....)))))))..))))))))...(((((....)))))))))))......-- ( -16.00, z-score = -0.92, R) >droSim1.chr3L 9905461 85 - 22553184 AUGCGCUCUGUUGCUUAACAUGCUGUUAUCGAUAAGCAUCUAUCGAAAUGUUAAGUUAUUUCGAAAGUUUGAAAAUGUAUUAAAA-- (((((((.(((((..((((.....)))).))))))))..((.((((((((......)))))))).)).........)))).....-- ( -17.40, z-score = -1.53, R) >consensus AUGCGCUCUGUUGCUUAACACUCUGUUAUCGAUAAGAAUGUAUCGAAAUGUUAAGCAAUUUCGAAAGUUUGAAAAUGAAU____A__ ............((((((((........((((((......))))))..))))))))..((((((....))))))............. (-14.28 = -13.28 + -1.00)
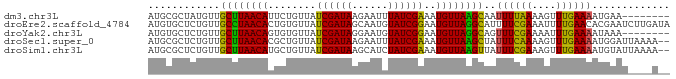
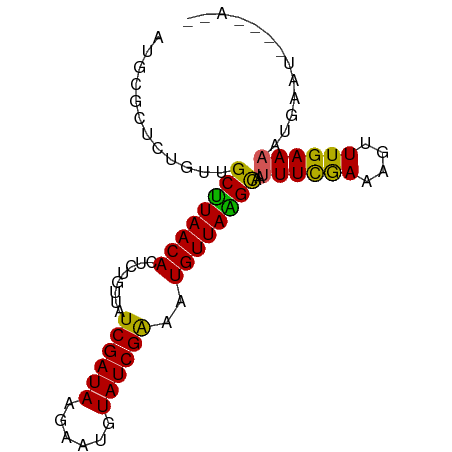
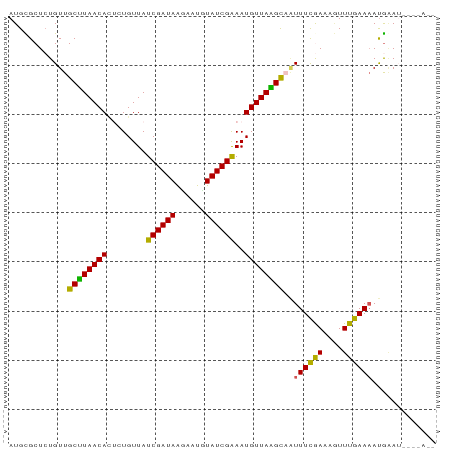
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:16:25 2011