
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 10,288,822 – 10,288,916 |
| Length | 94 |
| Max. P | 0.815157 |

| Location | 10,288,822 – 10,288,916 |
|---|---|
| Length | 94 |
| Sequences | 8 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 73.05 |
| Shannon entropy | 0.52665 |
| G+C content | 0.48918 |
| Mean single sequence MFE | -19.88 |
| Consensus MFE | -11.13 |
| Energy contribution | -11.84 |
| Covariance contribution | 0.70 |
| Combinations/Pair | 1.45 |
| Mean z-score | -1.26 |
| Structure conservation index | 0.56 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.78 |
| SVM RNA-class probability | 0.815157 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 10288822 94 + 24543557 UUAGAAAUACAAUAAAGUCGAGUCAAAGUGAAAAGUUUUACGCAGCGGCUGCCGCAGCUACUGCGUAUUUUCCGUCCAC--------ACCCCCCAAAAAGCC ...(((((((.................(((((....)))))((((..((((...))))..)))))))))))........--------............... ( -20.10, z-score = -1.80, R) >droSim1.chr3L 9678453 85 + 22553184 UUAGAAAUACAAUAAAGUCGAGUCAAAGUGAAAAGUUUUACGCAGCGGCUGCCGCAGCUACUGCGUAUUUUCC-----C------------CACCAAAAGCC ...(((((((.................(((((....)))))((((..((((...))))..)))))))))))..-----.------------........... ( -20.10, z-score = -1.94, R) >droSec1.super_0 2502930 102 + 21120651 UUAGAAAUACAAUAAAGUCGAGUCAAAGUGAAAAGUUUUACGCAGCGGCUGCCGCAGCUACUGCGUAUUUUCCGUCCACCCCGCCACACCCCACCAAAAGCC ...(((((((.................(((((....)))))((((..((((...))))..)))))))))))............................... ( -20.10, z-score = -1.28, R) >droYak2.chr3L 10277141 100 + 24197627 UUAGAAAUACAAUAAAGUCGAGUCAAAGUGAAAAGUUUUACGCAGCGGCUGCCGCAGCUACUGCGUAUUUUCCACCCACCCCCCU--CUACACCCAAAAGCC ................((.(((.....(((........(((((((..((((...))))..))))))).........)))....))--).))........... ( -22.33, z-score = -2.84, R) >droEre2.scaffold_4784 10281830 95 + 25762168 UUAGAAAUACAAUAAAGUCGAGUCAAAGUGAAAAGUUUUACGCAGCGGCUGCCGCAGCUACUGCGUAUUUUCCGCCCACCC-------CACCCCCAAAAGAC ...(((((((.................(((((....)))))((((..((((...))))..)))))))))))..........-------.............. ( -20.10, z-score = -1.93, R) >dp4.chrXR_group6 8682264 89 + 13314419 UUAGAAAUACAAUAAAGUCGAGUCAAAGUGAAAAGUUUUACGCAACGGCGGCGGCUGUUCCCCUCCCCACUUCGCC-CCCCCCUUUGGCC------------ .....................(((((((.(....(((......)))((.(((((..((..........)).)))))-))..)))))))).------------ ( -15.20, z-score = 1.08, R) >droPer1.super_27 1122326 90 + 1201981 UUAGAAAUACAAUAAAGUCGAGUCAAAGUGAAAAGUUUUACGCAACGGCGGCGGCUGUUCCCCUCCCUCCCCCUUCGCCCCCCCUUGGCC------------ ................((((((.....(((((..(((......)))((.((.((..(.....)..)).)).))))))).....)))))).------------ ( -16.00, z-score = 0.75, R) >droWil1.scaffold_181024 1429754 78 - 1751709 UUAGAAAUACAAUAAAGUGGAGUCAAAGUGAAAAGUUUUACGCAGUGGCGGCUGCCGCUGCUG-GUGCUACUACUCCAG----------------------- .................((((((....(((((....)))))(((.(((((((....)))))))-.)))....)))))).----------------------- ( -25.10, z-score = -2.12, R) >consensus UUAGAAAUACAAUAAAGUCGAGUCAAAGUGAAAAGUUUUACGCAGCGGCUGCCGCAGCUACUGCGUAUUUUCCGUCCACCC_______CC_C_CCAAAAGCC ......................................(((((((..((((...))))..)))))))................................... (-11.13 = -11.84 + 0.70)

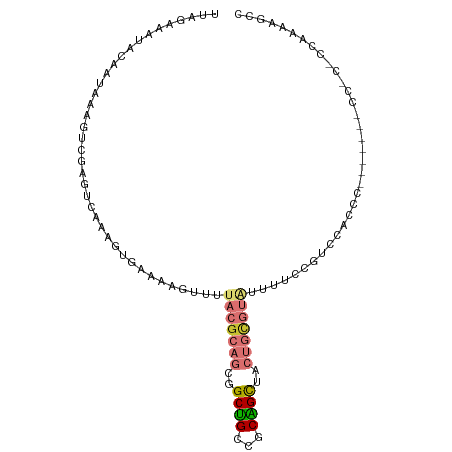

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:15:44 2011