
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 10,244,814 – 10,244,919 |
| Length | 105 |
| Max. P | 0.791187 |

| Location | 10,244,814 – 10,244,919 |
|---|---|
| Length | 105 |
| Sequences | 7 |
| Columns | 106 |
| Reading direction | forward |
| Mean pairwise identity | 85.64 |
| Shannon entropy | 0.28966 |
| G+C content | 0.48099 |
| Mean single sequence MFE | -33.26 |
| Consensus MFE | -20.06 |
| Energy contribution | -22.77 |
| Covariance contribution | 2.71 |
| Combinations/Pair | 1.10 |
| Mean z-score | -2.12 |
| Structure conservation index | 0.60 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.40 |
| SVM RNA-class probability | 0.681545 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
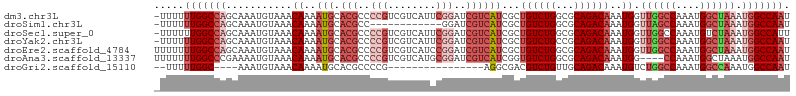
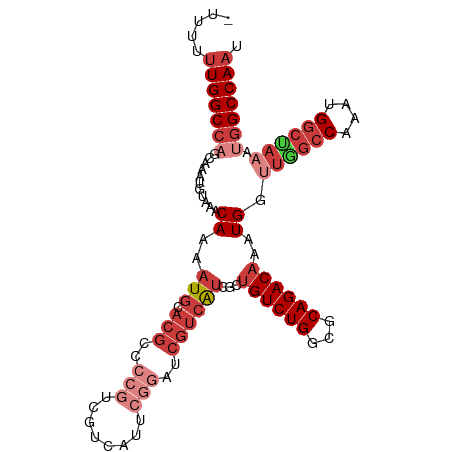
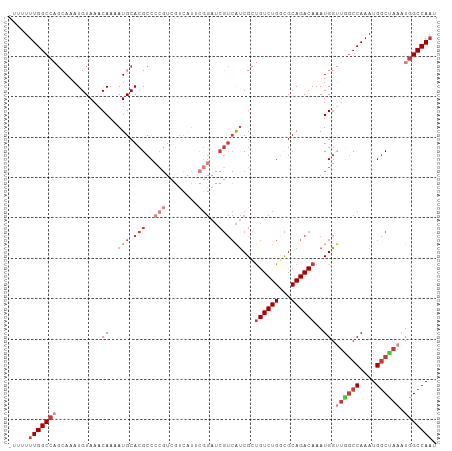
>dm3.chr3L 10244814 105 + 24543557 -UUUUUUGGCCAGCAAAUGUAAACAAAAUGCACGCCCCGUCGUCAUUCGGAUCGUCAUCGCUGUCUGGCGCAGACAAAUGGUUGGCCAAAUGGCUAAAUGGCCAAU -...((((((((((.............(((.(((..(((........)))..))))))...((((((...))))))....))))))))))(((((....))))).. ( -35.40, z-score = -2.12, R) >droSim1.chr3L 9634701 93 + 22553184 -UUUUUUGGCCAGCAAAUGUAAACAAAAUGCACGCC------------GGAUCGUCAUCGCUGUCUGGCGCAGACAAAUGGUUAGCCAAAUGGCUAAAUGGCCAAU -....((((((((((..((....))...))).((((------------((((.((....)).))))))))...........((((((....)))))).))))))). ( -34.00, z-score = -3.26, R) >droSec1.super_0 2459710 105 + 21120651 -UUUUUUGGCCAGCAAAUGUAAACAAAAUGCACGCCCCGUCGUCAUUCGGAUCGUCAUCGCUGUCUGGCGCAGACAAAUGGUUGGCCAAAUGUCUAAAUGGCCAUU -.....((((((((.............(((.(((..(((........)))..)))))).(((....)))))(((((..(((....)))..)))))...)))))).. ( -31.20, z-score = -1.36, R) >droYak2.chr3L 10232042 105 + 24197627 -UUUUUUGGCCAGCAAAUGUAAACAAAAUGCACGCCCCGUCGUCAUUCGGAUCGUCAUCGCUGUCUGCCGCAGACAAAUGGUUGGCCAAAUGGCUAAAUGGCCAAU -...((((((((((.............(((.(((..(((........)))..))))))...((((((...))))))....))))))))))(((((....))))).. ( -35.40, z-score = -2.38, R) >droEre2.scaffold_4784 10234466 106 + 25762168 UUUUUUUGGCCAGCAAAUGUAAACAAAAUGCACGCCCCGUCGUCAUCCGGAUCGUCAUCGCUGUCUGGCGCAGACAAAUGGUUGGCCAAAUGGCUAAAUGGCCAAU ....((((((((((.............(((.(((..(((........)))..))))))...((((((...))))))....))))))))))(((((....))))).. ( -34.90, z-score = -1.79, R) >droAna3.scaffold_13337 15159587 102 + 23293914 UUUUUUUGGCCCGAAAAUGUAAACAAAAUGCACGCCCCGUCGUCAUGCGGAUCGUCAUCGGUGUCUGGCGCAGACAAAUGG----CCAAAUGGCUAAAUGGCCAAU ....((((((((((...((....))...((.(((..((((......))))..)))))))).((((((...))))))...))----)))))(((((....))))).. ( -31.40, z-score = -0.96, R) >droGri2.scaffold_15110 11925330 84 - 24565398 --UUUUUGGC----AAAUGUAAACAAAAUGCACGCCCCG----------------AGGCGACGUCUGUUGCAGACAAAUGUCUGGCCAAAUGGCCAAAUGGCCAAU --..((((((----.......((((..(((..((((...----------------.)))).))).)))).(((((....)))))))))))(((((....))))).. ( -30.50, z-score = -2.96, R) >consensus _UUUUUUGGCCAGCAAAUGUAAACAAAAUGCACGCCCCGUCGUCAUUCGGAUCGUCAUCGCUGUCUGGCGCAGACAAAUGGUUGGCCAAAUGGCUAAAUGGCCAAU .....(((((((...........((..(((.(((..(((........)))..))))))...((((((...))))))..)).((((((....)))))).))))))). (-20.06 = -22.77 + 2.71)
| Location | 10,244,814 – 10,244,919 |
|---|---|
| Length | 105 |
| Sequences | 7 |
| Columns | 106 |
| Reading direction | reverse |
| Mean pairwise identity | 85.64 |
| Shannon entropy | 0.28966 |
| G+C content | 0.48099 |
| Mean single sequence MFE | -33.06 |
| Consensus MFE | -22.22 |
| Energy contribution | -24.63 |
| Covariance contribution | 2.40 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.01 |
| Structure conservation index | 0.67 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.70 |
| SVM RNA-class probability | 0.791187 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
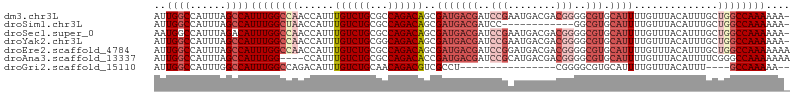
>dm3.chr3L 10244814 105 - 24543557 AUUGGCCAUUUAGCCAUUUGGCCAACCAUUUGUCUGCGCCAGACAGCGAUGACGAUCCGAAUGACGACGGGGCGUGCAUUUUGUUUACAUUUGCUGGCCAAAAAA- ..((((......))))((((((((..((((((((((...))))))..))))(((..(((........)))..)))(((...((....))..)))))))))))...- ( -34.10, z-score = -1.81, R) >droSim1.chr3L 9634701 93 - 22553184 AUUGGCCAUUUAGCCAUUUGGCUAACCAUUUGUCUGCGCCAGACAGCGAUGACGAUCC------------GGCGUGCAUUUUGUUUACAUUUGCUGGCCAAAAAA- .(((((((.((((((....))))))..........(((((.((...((....)).)).------------)))))(((...((....))..))))))))))....- ( -30.70, z-score = -2.39, R) >droSec1.super_0 2459710 105 - 21120651 AAUGGCCAUUUAGACAUUUGGCCAACCAUUUGUCUGCGCCAGACAGCGAUGACGAUCCGAAUGACGACGGGGCGUGCAUUUUGUUUACAUUUGCUGGCCAAAAAA- ..((((((..((((((..(((....)))..))))))....((((((.(((((((..(((........)))..))).))))))))))........)))))).....- ( -34.50, z-score = -2.33, R) >droYak2.chr3L 10232042 105 - 24197627 AUUGGCCAUUUAGCCAUUUGGCCAACCAUUUGUCUGCGGCAGACAGCGAUGACGAUCCGAAUGACGACGGGGCGUGCAUUUUGUUUACAUUUGCUGGCCAAAAAA- ..((((......))))((((((((..((((((((((...))))))..))))(((..(((........)))..)))(((...((....))..)))))))))))...- ( -34.10, z-score = -1.62, R) >droEre2.scaffold_4784 10234466 106 - 25762168 AUUGGCCAUUUAGCCAUUUGGCCAACCAUUUGUCUGCGCCAGACAGCGAUGACGAUCCGGAUGACGACGGGGCGUGCAUUUUGUUUACAUUUGCUGGCCAAAAAAA ..((((......))))((((((((..((((((((((...))))))..))))(((..(((........)))..)))(((...((....))..))))))))))).... ( -35.40, z-score = -1.83, R) >droAna3.scaffold_13337 15159587 102 - 23293914 AUUGGCCAUUUAGCCAUUUGG----CCAUUUGUCUGCGCCAGACACCGAUGACGAUCCGCAUGACGACGGGGCGUGCAUUUUGUUUACAUUUUCGGGCCAAAAAAA ..((((......))))(((((----((...((((((...))))))..(((((((..(((........)))..))).))))...............))))))).... ( -31.40, z-score = -1.17, R) >droGri2.scaffold_15110 11925330 84 + 24565398 AUUGGCCAUUUGGCCAUUUGGCCAGACAUUUGUCUGCAACAGACGUCGCCU----------------CGGGGCGUGCAUUUUGUUUACAUUU----GCCAAAAA-- ..(((((....)))))(((((((((((....))))).((((((.((((((.----------------...)))).))..)))))).......----))))))..-- ( -31.20, z-score = -2.91, R) >consensus AUUGGCCAUUUAGCCAUUUGGCCAACCAUUUGUCUGCGCCAGACAGCGAUGACGAUCCGAAUGACGACGGGGCGUGCAUUUUGUUUACAUUUGCUGGCCAAAAAA_ ..((((......))))((((((((......((((((...))))))..(((((((.((((........)))).))).))))..............)))))))).... (-22.22 = -24.63 + 2.40)


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:15:30 2011