
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 10,078,242 – 10,078,353 |
| Length | 111 |
| Max. P | 0.500000 |

| Location | 10,078,242 – 10,078,353 |
|---|---|
| Length | 111 |
| Sequences | 9 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 79.12 |
| Shannon entropy | 0.41210 |
| G+C content | 0.41852 |
| Mean single sequence MFE | -27.06 |
| Consensus MFE | -12.63 |
| Energy contribution | -12.68 |
| Covariance contribution | 0.05 |
| Combinations/Pair | 1.22 |
| Mean z-score | -1.84 |
| Structure conservation index | 0.47 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.00 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
>dm3.chr3L 10078242 111 + 24543557 AUCCCAUUGCUGUUAAUGUUUACGAUUGUCGUUUUCAAUUUCAGUUAAUUGAAAC-------AAGGGUAACACACAGCAACGCAUUAAGCCCGCU-CCAAAGGGACCCUCACUUCUCCC- .((((.(((((((...((((.((..((((....(((((((......)))))))))-------))..)))))).))))))).((.....)).....-.....))))..............- ( -24.70, z-score = -1.95, R) >droSec1.super_0 2293484 111 + 21120651 AUCCCAUUGCUGUUAAUGUUUACGAUUGUCGUUUUCAAUUUCAGUUAAUUGAAAC-------AAGGGCAACAAACAGCAAGGCAUUAAGCCCGCU-CCAAAGGGACCUUCAAUUCUCCC- .((((.((((((((..((((((((.....)))((((((((......)))))))).-------..)))))...))))))))(((.....)))....-.....))))..............- ( -28.00, z-score = -2.48, R) >droSim1.chr3L 9458434 111 + 22553184 AUCCCAUUGCUGUUAAUGUUUACGAUUGUCGUUUUCAAUUUCAGUUAAUUGAAAC-------AAGGGCAACAAACAGCAAGGCAUUAAGCCCGCU-CCAAAGGGACCUUCAAUUCUCCC- .((((.((((((((..((((((((.....)))((((((((......)))))))).-------..)))))...))))))))(((.....)))....-.....))))..............- ( -28.00, z-score = -2.48, R) >droYak2.chr3L 10056700 107 + 24197627 AUCCCAUUGCUGUUAAUGUUUACGAUUGUCGUUUUCAAUUUCAGUUAAUUGAAAC-------AAGGGCAACA----GCAAUGCAUUAAGCCCGCU-CCAAAGGGACCUUCAAUUCUCCC- ....(((((((((...((((((((.....)))((((((((......)))))))).-------..))))))))----)))))).......(((...-.....)))...............- ( -24.20, z-score = -1.64, R) >droEre2.scaffold_4784 10057688 107 + 25762168 AUCCCAUUGCUGUUAAUGUUUACGAUUGUCGUUUUCAAUUUCAGUUAAUUGAAAC-------AAGGGCAACA----GCAACGCAUUAAGCCAGCU-CCAAAGGGACCUUCAAUUCUCCC- .((((...(((((((((((((.(..(((((.(((((((((......)))))))..-------)).)))))..----).)).)))))))..)))).-.....))))..............- ( -23.80, z-score = -1.67, R) >droAna3.scaffold_13337 14989744 108 + 23293914 UUCCCAUUGCUGUUAAUGUUUACGAUUGUCGUUUUCAAUUUCAGUUAAUUGAAAC-------AAGGGCAGCC----CCAUCACAUUAAGCUUCUUGCCAAAGGGACCUGCAAUUCUCAC- .((((...((((((..(((((.((((((.(.............).))))))))))-------)..)))))).----............((.....))....))))..............- ( -21.72, z-score = -0.66, R) >droPer1.super_27 896794 118 + 1201981 AUCGCACUGCUGUUAAUGGUUACGAUUGUCGUUUUCAAUUUCAGUUAAUUGAAACUGCUGGCAAGGGUAACA-ACACCAUGACACAGUGCUCCCU-UUUGAGGGACCCUCUCUGGCUCCA ...((((((.(((((.(((((((..(((((((((((((((......))))))))..)).)))))..)))...-..)))))))))))))))(((((-....))))).((.....))..... ( -36.70, z-score = -2.40, R) >dp4.chrXR_group6 5752891 118 - 13314419 AUCGCACUGCCGUUAAUGGUUACGAUUGUCGUUUUCAAUUUCAGUUAAUUGAAACUGCUGGCAAGGGUAACA-ACACCAUGACACAGUGCUCCCU-UUUGAGGGACCCUCUCUGGCUCCA ...((((((..((((.(((((((..(((((((((((((((......))))))))..)).)))))..)))...-..)))))))).))))))(((((-....))))).((.....))..... ( -34.50, z-score = -1.75, R) >droWil1.scaffold_181024 1019414 108 - 1751709 AUCUCAUAGCUGUUAAUGUUUACGAUUGUCGUUUUCAAUUUCAGUUAAUUGAAAUGGCUCA-AAAGGCAACC-----AAGAACAUUAUGCUCC------AGGGAAUUUUCAUUCACUUCC .((((..(((...((((((((......(((((((.(((((......))))))))))))...-...(....).-----..)))))))).)))..------.))))................ ( -21.90, z-score = -1.50, R) >consensus AUCCCAUUGCUGUUAAUGUUUACGAUUGUCGUUUUCAAUUUCAGUUAAUUGAAAC_______AAGGGCAACA_ACAGCAAGGCAUUAAGCCCGCU_CCAAAGGGACCUUCAAUUCUCCC_ .((((...((((..(.((...(((.....)))...)).)..))))...................((((....................)))).........))))............... (-12.63 = -12.68 + 0.05)
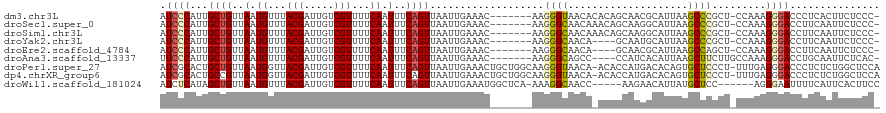

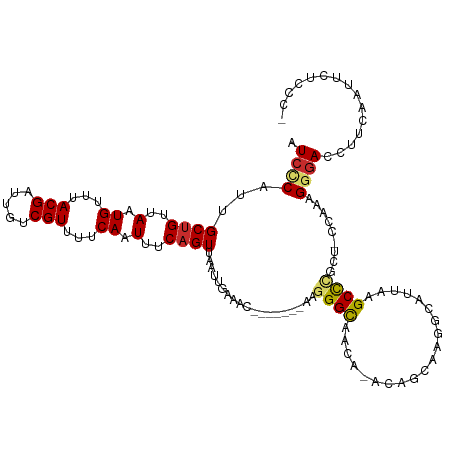
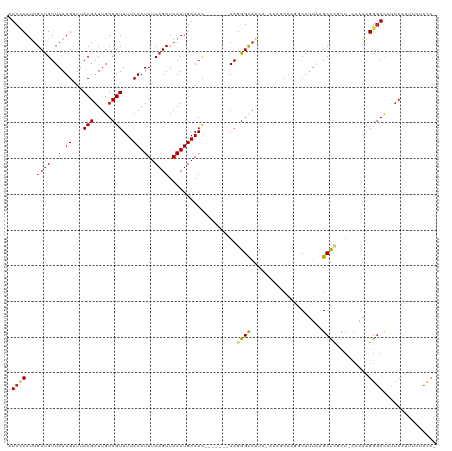
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:15:01 2011