| Sequence ID | dm3.chr2L |
|---|---|
| Location | 4,714,564 – 4,714,654 |
| Length | 90 |
| Max. P | 0.835319 |

| Location | 4,714,564 – 4,714,654 |
|---|---|
| Length | 90 |
| Sequences | 10 |
| Columns | 95 |
| Reading direction | forward |
| Mean pairwise identity | 69.02 |
| Shannon entropy | 0.65236 |
| G+C content | 0.50052 |
| Mean single sequence MFE | -19.77 |
| Consensus MFE | -11.86 |
| Energy contribution | -11.23 |
| Covariance contribution | -0.63 |
| Combinations/Pair | 1.50 |
| Mean z-score | -0.56 |
| Structure conservation index | 0.60 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.85 |
| SVM RNA-class probability | 0.835319 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
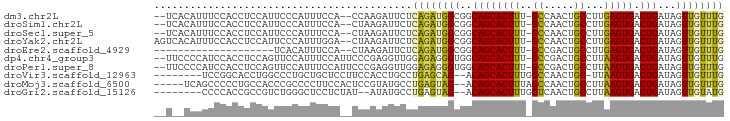
>dm3.chr2L 4714564 90 + 23011544 --UCACAUUUCCACCUCCAUUCCCAUUUCCA--CCAAGAUUCUCAGAUGGCGGCAGCACUUU-GCCAACUGCCUUGAGUGACUGAUAGUUGUUUG --.............................--..........((((..((..((((((((.-((.....))...))))).)))...))..)))) ( -16.40, z-score = -0.19, R) >droSim1.chr2L 4583002 90 + 22036055 --UCACAUUUCCACCUCCAUUCCCAUUUCCA--CUAAGAUUCUCAGAUGGCGGCAGCACUUU-GCCAACUGCCUUGAGUGACUGAUAGUUGUUUG --.............................--..........((((..((..((((((((.-((.....))...))))).)))...))..)))) ( -16.40, z-score = -0.21, R) >droSec1.super_5 2790989 90 + 5866729 --UCACAUUUCCACCUCCAUUCCCAUUUCCA--CUAAGAUUCUCAGAUGGCGGCAGCACUUU-GCCAACUGCCUUGAGUGACUGAUAGUUGUUUG --.............................--..........((((..((..((((((((.-((.....))...))))).)))...))..)))) ( -16.40, z-score = -0.21, R) >droYak2.chr2L 4721987 92 + 22324452 AGUCACAUUUCCACCUCCAUUCCCAUUUGGA--CUAAGAUUCUCAGAUGGCGGCAGCACUUU-GCCAACUGCCUUGAGUGACUGAUAGUUGUUUG ((((((................(((((((((--.......)).))))))).(((((......-.....)))))....))))))............ ( -22.80, z-score = -1.08, R) >droEre2.scaffold_4929 4789174 72 + 26641161 --------------------UCACAUUUCCA--CUAAGAUUCUCAGAUGGCGGCAGCACUUU-GCCGACUGCCUUGAGUGACUGAUAGUUGUUUG --------------------...........--..........((((..((..((((((((.-((.....))...))))).)))...))..)))) ( -16.40, z-score = -0.29, R) >dp4.chr4_group3 560689 92 - 11692001 --UUCCCCAUCCACCUCCAGUUCCAUUUCCAUUCCCGAGGUUGGAGAGGGUGGCAGCACUUU-GCCGACUGCCUUAAGUGACUGAUAGUUGUUUG --..(((..((((((((...................)))).))))..))).((((((.....-...).))))).((((..((.....))..)))) ( -24.91, z-score = -0.83, R) >droPer1.super_8 1604582 92 - 3966273 --UUCCCCAUCCACCUCCAGUUCCAUUUCCAUUCCCGAGGUUGGAGAGGGUGGCAGCACUUU-GCCGACUGCCUUAAGUGACUGAUAGUUGUUUG --..(((..((((((((...................)))).))))..))).((((((.....-...).))))).((((..((.....))..)))) ( -24.91, z-score = -0.83, R) >droVir3.scaffold_12963 8630163 84 - 20206255 --------UCCGGCACCUGGCCCUGCUGCUCCUUCCACCUGCCUGAGCAG--ACAGCACUUUGGCCAACUGC-UUAAGUGACUGAUAGUUGUUUG --------...((((..((((((((((((((.............))))))--.)))......)))))..)))-)((((..((.....))..)))) ( -25.82, z-score = -1.36, R) >droMoj3.scaffold_6500 12090850 88 - 32352404 -----UCAGCCCCCUGCCACCCGCCCCUUCCACUCCGUAUGCCUGAGUAG--ACAGCACUUUAGCCAACUGCCUUAAGUGACUGAUAGUUGUUUG -----.((((..................((.((((.(.....).)))).)--)((((((((..((.....))...))))).)))...)))).... ( -14.50, z-score = -0.45, R) >droGri2.scaffold_15126 3255239 83 - 8399593 --------CCCCACCGCCGUCUGGGCUCCUCUAU--AUAUGCCUGAGUAG--ACAGCACUUUGGUCAACUGCCUUAAGUGACUGAUAGUUGUAUG --------.......((.((((((((........--....))))....))--)).))((((..((((.((......))))))..).)))...... ( -19.20, z-score = -0.17, R) >consensus __U__CAUUUCCACCUCCAUUCCCAUUUCCA__CCAAGAUUCUCAGAUGGCGGCAGCACUUU_GCCAACUGCCUUAAGUGACUGAUAGUUGUUUG ...........................................((((((((..((((((((..((.....))...))))).)))...)))))))) (-11.86 = -11.23 + -0.63)


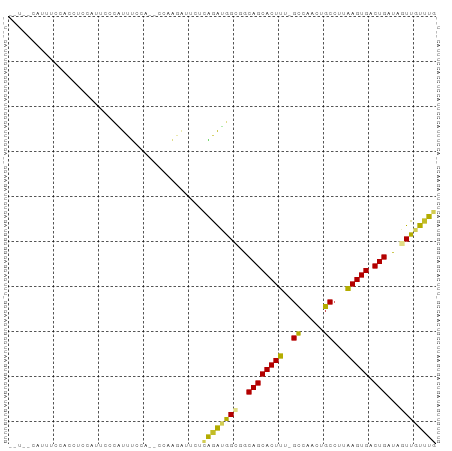
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:17:15 2011