
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 9,714,878 – 9,714,978 |
| Length | 100 |
| Max. P | 0.539853 |

| Location | 9,714,878 – 9,714,978 |
|---|---|
| Length | 100 |
| Sequences | 7 |
| Columns | 109 |
| Reading direction | forward |
| Mean pairwise identity | 66.95 |
| Shannon entropy | 0.64284 |
| G+C content | 0.48553 |
| Mean single sequence MFE | -28.46 |
| Consensus MFE | -11.07 |
| Energy contribution | -12.20 |
| Covariance contribution | 1.13 |
| Combinations/Pair | 1.50 |
| Mean z-score | -1.19 |
| Structure conservation index | 0.39 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.09 |
| SVM RNA-class probability | 0.539853 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
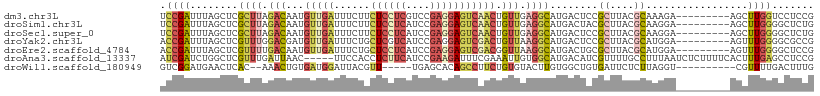
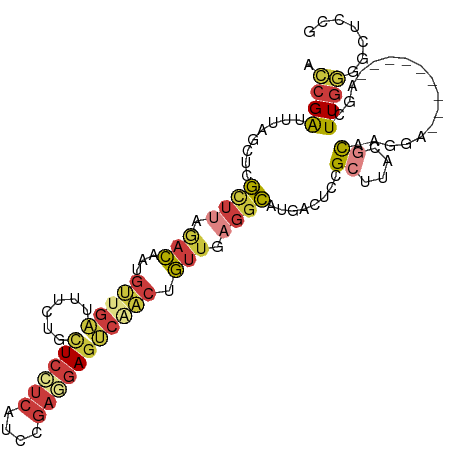
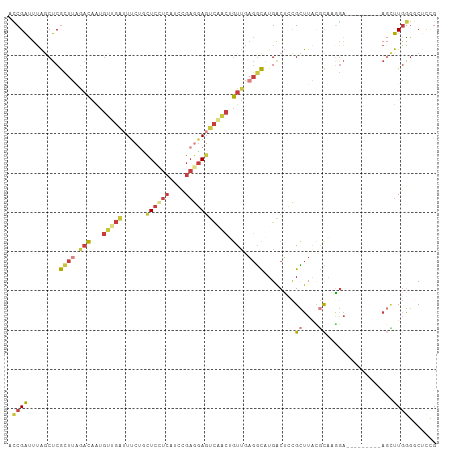
>dm3.chr3L 9714878 100 + 24543557 UCCGAUUUAGCUCGCUUAGACAAUGUUGAUUUCUUCUCCUCGUCCGAGGAGUCAACUGUUGAGGCAUGACUCCGCUUACGCAAAGA---------AGCUUGGUCCUCCG ...(((..(((((((((.((((..(((((......((((((....))))))))))))))).))))........((....))...).---------))))..)))..... ( -28.80, z-score = -1.30, R) >droSim1.chr3L 9095470 100 + 22553184 UCCGAUUUAGCUCGCUUAGACAAUGUUGAUUUCUUCUCCUCAUCCGAGGAGUCAACUGUUGAGGCAUGACUACGCUUACGCAAGGA---------AGCUUGGGGCUCUG ........(((((((((.((((..(((((......((((((....))))))))))))))).))))........((((.(....).)---------)))...)))))... ( -32.00, z-score = -1.73, R) >droSec1.super_0 1948512 100 + 21120651 UCCGAUUUAGCUCGCUUAGACAAUGUUGAUUUCUUCUCCUCAUCCGAGGAGUCAACUGUUGAGGCAUGACUCCGCUUACGCAAGGA---------AGCUUGGGGCUCUG ........(((((((((.((((..(((((......((((((....))))))))))))))).))))........((((.(....).)---------)))...)))))... ( -32.00, z-score = -1.41, R) >droYak2.chr3L 21979758 100 - 24197627 ACCGAUUUAGCUCGUUUGGACGAUGUUGAUUUCUGCUCGUCAUCCGAGGAGUCGACUGUUAAGGCAUGACUCCGCUUACGCAUGGA---------AGUUUGGGGCGCCG .........((((.....(((((.((.(....).))))))).((((.(((((((..(((....))))))))))((....)).))))---------......)))).... ( -29.70, z-score = 0.06, R) >droEre2.scaffold_4784 9701969 100 + 25762168 ACCGAUUUAGCUCGUUUUGACAAUGUUGAUUUCUGCUCCUCAUCCGAGGAGUCGACGGUUAAGGCAUGACUGCGCUUACGCAUGGA---------AGUUUGGGGCUCCG ........(((((((((((((...((((......(((((((....))))))))))).)))))))).....((((....))))....---------......)))))... ( -29.70, z-score = -0.57, R) >droAna3.scaffold_13337 19630062 104 - 23293914 AUCGAUCUGGCUCGUUUGAUUAAC-----UUCCACCUCUUCAUCCGAAGAUUUCGAAAUUGUGGCAUGACAUCGUUUUGCCUUUAAUCUCUUUUCACUUUGAGCCUCCG ........((((((..(((.....-----.......(((((....)))))....((.((((.((((.(((...))).))))..)))).))...)))...)))))).... ( -23.00, z-score = -2.42, R) >droWil1.scaffold_180949 5166872 92 - 6375548 GUCGGAUGAACUCAC--AAACUGUGAUGGAUUACGUU-----UGAGCACAGCCUUCUGUGUACUUGUGGCUGUGAUUCUCUUAGGU----------CGUUUUGACUUUG (((((((((.((..(--((((.((((....)))))))-----))..(((((((....(....)....)))))))........)).)----------)).)))))).... ( -24.00, z-score = -0.94, R) >consensus ACCGAUUUAGCUCGCUUAGACAAUGUUGAUUUCUGCUCCUCAUCCGAGGAGUCAACUGUUGAGGCAUGACUCCGCUUACGCAAGGA_________AGCUUGGGGCUCCG .((((........((((.(((...(((((......((((((....))))))))))).))).))))........((....)).................))))....... (-11.07 = -12.20 + 1.13)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:14:14 2011