
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 9,164,776 – 9,164,868 |
| Length | 92 |
| Max. P | 0.583869 |

| Location | 9,164,776 – 9,164,868 |
|---|---|
| Length | 92 |
| Sequences | 10 |
| Columns | 103 |
| Reading direction | forward |
| Mean pairwise identity | 83.26 |
| Shannon entropy | 0.33155 |
| G+C content | 0.43442 |
| Mean single sequence MFE | -22.96 |
| Consensus MFE | -16.21 |
| Energy contribution | -15.84 |
| Covariance contribution | -0.37 |
| Combinations/Pair | 1.22 |
| Mean z-score | -1.33 |
| Structure conservation index | 0.71 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.04 |
| SVM RNA-class probability | 0.512964 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 9164776 92 + 24543557 CCAGUCGCCAGGCACUUCAAUGCCU--GUUUAUUUGCGUCUAUUCUACGCUAAACUGUGCUAUUAAAGCGCGUUUGUUGUUAUUCCAACUCUGC--------- ..((.(((((((((......)))))--).......))).)).....(((.(((((.(((((.....)))))))))).)))..............--------- ( -21.81, z-score = -1.84, R) >droSim1.chr3L 8562406 92 + 22553184 GCAGUCGCCAGGCACUUCAAUGCCU--GUUUAUUUGCGUCUAUUCUACGCUAAACUGUGCUAUUAAAGCGCGUUUGUUGUUAUUCCAACUCUGC--------- ((((....((((((......)))))--).......((((.......)))).((((.(((((.....)))))))))((((......)))).))))--------- ( -24.10, z-score = -2.16, R) >droSec1.super_0 1422079 92 + 21120651 GCAGUCGCCAGGCACUUCAAUGCCU--GUUUAUUUGCGUCUAUUCUACGCUAAACUGUGCUAUUAAAGCGCGUUUGUUGUUAUUCCAACUCUGC--------- ((((....((((((......)))))--).......((((.......)))).((((.(((((.....)))))))))((((......)))).))))--------- ( -24.10, z-score = -2.16, R) >droYak2.chr3L 21440664 92 - 24197627 GCAGUCGCCAGGCACUUCAAUGCCU--GUUUAUUUGCGACUAUUCUACGCUAAACUGUGCUAUUAAAGCGCGUUUGUUGUUAUUCCAGCUCUGC--------- ((((((((((((((......)))))--).......)))))).....(((.(((((.(((((.....)))))))))).))).......)).....--------- ( -27.61, z-score = -2.53, R) >droEre2.scaffold_4784 20879110 92 - 25762168 GCAGUCGCCAGGCACUUCAAUGCCU--GUUUAUUUGCGUCUAUUCUACGCUAAACUGUGCUAUUAAAGCGCGUUUGUUGUUAUUCCAGCUCGGC--------- ((((.(((((((((......)))))--).......))).)).....(((.(((((.(((((.....)))))))))).))).......)).....--------- ( -23.71, z-score = -0.93, R) >droAna3.scaffold_13337 19564203 100 + 23293914 CCAGGCGA-AGGCACUUCAAUGCCU--GUUUAUUUGCGUCUAUUCUACGCUAAACUGUGCUAUUAAAGCGCGUUUGUUGUUAUUCCAACUCUGCCAGAAGCCA ...(((..-(((((......)))))--(((((...((((.......))))))))).(((((.....)))))((..((((......))))...)).....))). ( -24.90, z-score = -0.99, R) >droWil1.scaffold_181024 340083 89 - 1751709 GAGGUAG--AGGCACUUCAAUGCCU--GUUUAUUUGCGUCUAUUCUACGCUAAACUGUGCUAUUAAAGCGCGUUUGUUGUUAUUUCAACUGUC---------- (((((((--(((((......)))))--........((((.......))))(((((.(((((.....))))))))))...))))))).......---------- ( -21.60, z-score = -1.30, R) >droVir3.scaffold_13049 748268 86 + 25233164 --------AAGGCACAACCAUGACUGUGUUUAUUUGCGUCUAUUCUGCGCUAAACUGUGCUAUUAAAGCGCGUUUGUUGGCAUUUUAACUGUCU--------- --------.(((((((........)))))))....((((.......))))(((((.(((((.....))))))))))..((((.......)))).--------- ( -18.80, z-score = -0.09, R) >droMoj3.scaffold_6680 24268536 94 - 24764193 UUUAUGGCAAGUCCCUGGCAUGACUCUGUUUAUUUGCGUCUAUUCUGCGCUAAACUGUGCUAUUAAAGCGCGUUUGUUGGCAUUUUAACUGUCU--------- .....((((.((..(..(((.(((...(((((...((((.......))))))))).(((((.....)))))))))))..).......)))))).--------- ( -18.70, z-score = 0.27, R) >droGri2.scaffold_15110 1971441 86 + 24565398 --------AAGACGCAAGCAUGACUGUGUUUAUUUGCGUCUAUUCUGCGCUAAACUGUGCUAUUAAAGCGCGUUUGUUGGCAUUUUAACUGUCU--------- --------.((((((((((((....))))....)))))))).......(((((..((((((.....))))))....))))).............--------- ( -24.30, z-score = -1.56, R) >consensus GCAGUCGCCAGGCACUUCAAUGCCU__GUUUAUUUGCGUCUAUUCUACGCUAAACUGUGCUAUUAAAGCGCGUUUGUUGUUAUUCCAACUCUGC_________ .........(((((......)))))..(((((...((((.......))))))))).(((((.....)))))....((((......)))).............. (-16.21 = -15.84 + -0.37)


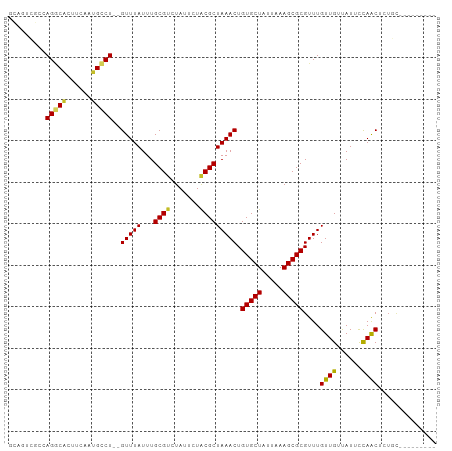
| Location | 9,164,776 – 9,164,868 |
|---|---|
| Length | 92 |
| Sequences | 10 |
| Columns | 103 |
| Reading direction | reverse |
| Mean pairwise identity | 83.26 |
| Shannon entropy | 0.33155 |
| G+C content | 0.43442 |
| Mean single sequence MFE | -24.76 |
| Consensus MFE | -12.66 |
| Energy contribution | -13.02 |
| Covariance contribution | 0.36 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.16 |
| Structure conservation index | 0.51 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.19 |
| SVM RNA-class probability | 0.583869 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 9164776 92 - 24543557 ---------GCAGAGUUGGAAUAACAACAAACGCGCUUUAAUAGCACAGUUUAGCGUAGAAUAGACGCAAAUAAAC--AGGCAUUGAAGUGCCUGGCGACUGG ---------.(((.((((......))))...((((((.....)))...((((.((((.......))))))))...(--((((((....)))))))))).))). ( -29.50, z-score = -3.81, R) >droSim1.chr3L 8562406 92 - 22553184 ---------GCAGAGUUGGAAUAACAACAAACGCGCUUUAAUAGCACAGUUUAGCGUAGAAUAGACGCAAAUAAAC--AGGCAUUGAAGUGCCUGGCGACUGC ---------((((.((((......))))...((((((.....)))...((((.((((.......))))))))...(--((((((....)))))))))).)))) ( -31.40, z-score = -4.26, R) >droSec1.super_0 1422079 92 - 21120651 ---------GCAGAGUUGGAAUAACAACAAACGCGCUUUAAUAGCACAGUUUAGCGUAGAAUAGACGCAAAUAAAC--AGGCAUUGAAGUGCCUGGCGACUGC ---------((((.((((......))))...((((((.....)))...((((.((((.......))))))))...(--((((((....)))))))))).)))) ( -31.40, z-score = -4.26, R) >droYak2.chr3L 21440664 92 + 24197627 ---------GCAGAGCUGGAAUAACAACAAACGCGCUUUAAUAGCACAGUUUAGCGUAGAAUAGUCGCAAAUAAAC--AGGCAUUGAAGUGCCUGGCGACUGC ---------((.(((((((.....).........(((.....))).)))))).))......(((((((.......(--((((((....)))))))))))))). ( -28.31, z-score = -2.35, R) >droEre2.scaffold_4784 20879110 92 + 25762168 ---------GCCGAGCUGGAAUAACAACAAACGCGCUUUAAUAGCACAGUUUAGCGUAGAAUAGACGCAAAUAAAC--AGGCAUUGAAGUGCCUGGCGACUGC ---------((((.(.((......)).)....((((((((((.((...(((((((((.......))))...)))))--..)))))))))))).))))...... ( -26.00, z-score = -1.79, R) >droAna3.scaffold_13337 19564203 100 - 23293914 UGGCUUCUGGCAGAGUUGGAAUAACAACAAACGCGCUUUAAUAGCACAGUUUAGCGUAGAAUAGACGCAAAUAAAC--AGGCAUUGAAGUGCCU-UCGCCUGG ........(((...((((......))))....((((((((((.((...(((((((((.......))))...)))))--..))))))))))))..-..)))... ( -29.90, z-score = -2.03, R) >droWil1.scaffold_181024 340083 89 + 1751709 ----------GACAGUUGAAAUAACAACAAACGCGCUUUAAUAGCACAGUUUAGCGUAGAAUAGACGCAAAUAAAC--AGGCAUUGAAGUGCCU--CUACCUC ----------....((((......))))....((((((((((.((...(((((((((.......))))...)))))--..))))))))))))..--....... ( -23.30, z-score = -3.35, R) >droVir3.scaffold_13049 748268 86 - 25233164 ---------AGACAGUUAAAAUGCCAACAAACGCGCUUUAAUAGCACAGUUUAGCGCAGAAUAGACGCAAAUAAACACAGUCAUGGUUGUGCCUU-------- ---------.............((((((....(((((..(((......))).)))))......(((.............)))...)))).))...-------- ( -15.92, z-score = 0.12, R) >droMoj3.scaffold_6680 24268536 94 + 24764193 ---------AGACAGUUAAAAUGCCAACAAACGCGCUUUAAUAGCACAGUUUAGCGCAGAAUAGACGCAAAUAAACAGAGUCAUGCCAGGGACUUGCCAUAAA ---------.....((((((.(((........))).)))))).((...(((((((((......).)))...))))).(((((........)))))))...... ( -15.10, z-score = 0.39, R) >droGri2.scaffold_15110 1971441 86 - 24565398 ---------AGACAGUUAAAAUGCCAACAAACGCGCUUUAAUAGCACAGUUUAGCGCAGAAUAGACGCAAAUAAACACAGUCAUGCUUGCGUCUU-------- ---------.......................(((((..(((......))).))))).....((((((((................)))))))).-------- ( -16.79, z-score = -0.21, R) >consensus _________GCAGAGUUGGAAUAACAACAAACGCGCUUUAAUAGCACAGUUUAGCGUAGAAUAGACGCAAAUAAAC__AGGCAUUGAAGUGCCUGGCGACUGC ................................(((((.....)))...(((((........)))))))..........((((((....))))))......... (-12.66 = -13.02 + 0.36)



Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:13:08 2011