
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 8,282,963 – 8,283,053 |
| Length | 90 |
| Max. P | 0.807912 |

| Location | 8,282,963 – 8,283,053 |
|---|---|
| Length | 90 |
| Sequences | 5 |
| Columns | 115 |
| Reading direction | forward |
| Mean pairwise identity | 67.38 |
| Shannon entropy | 0.56533 |
| G+C content | 0.44653 |
| Mean single sequence MFE | -27.86 |
| Consensus MFE | -11.62 |
| Energy contribution | -11.86 |
| Covariance contribution | 0.24 |
| Combinations/Pair | 1.39 |
| Mean z-score | -1.38 |
| Structure conservation index | 0.42 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.12 |
| SVM RNA-class probability | 0.550730 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 8282963 90 + 24543557 AUGGUCUGACGACCGACGGUUCCUUCGAUAAGUCGUUACGCUAUCGAUGUCUUACUCUCGAUAGUUGUCAGAAGUGCCUUUUUCUUCUUC------------------------- ....((((((((((((.(((....((((((.((......))))))))......))).)))...)))))))))..................------------------------- ( -22.40, z-score = -0.83, R) >droSim1.chr3L 7742235 104 + 22553184 AUGGCCUGGCGACCGACGGUUCCUUCGAUAAGUCGUUAAGUUAUCGAUGUCUUACUUUCGAUAGUUGGCGAUAGUGUCUUUUUGCUAGUUUUUGUUUCUCCAUC----------- .....(((((((..((.(....).))((((.((((((((.(((((((..........)))))))))))))))..))))...)))))))................----------- ( -25.10, z-score = -0.97, R) >droSec1.super_0 563775 104 + 21120651 AUGGCCUGGCGACCGACGGUUCCUUCGAUAAGUCCUUAAGUUAUCGAUGUCUUACUUUCGAUAGUUGGCGAUAGUGUCUUUUAGCCAGGUUAAGUUUCUCCUCC----------- .(((((((((...(((.(((....(((((((.(.....).)))))))......))).))).(((..((((....))))..))))))))))))............----------- ( -29.40, z-score = -1.71, R) >droYak2.chr3L 20581374 114 - 24197627 AUGGGCUGGCCACCCUCGAUAGUUUCGAUACAUUAACGAGCUAUCGA-AUUUUACUAUCGAUAGUUGGCGGAAGUGACUUUUUGCGAAGUUACGUUAAAUUGGAGAUUCUUCUUC ...(((..(((((..((((((((((((((((........).))))))-)....))))))))..).))))....((((((((....))))))))))).....(((((....))))) ( -32.10, z-score = -2.08, R) >droEre2.scaffold_4784 20008308 102 - 25762168 AUGGGCUGACGGCCGUCGAUAGUUUCGAUAAGUUGACGAGCUAUCGAUGUUGUACUUUCGAUAGUUGGCGGAAGUGCCUUCUUGCGAACUUUCUUUUUCCUC------------- ..(((...(((((.((((((((((((((....))))..)))))))))))))))......((.((((.(((((((...)))).))).)))).))....)))..------------- ( -30.30, z-score = -1.29, R) >consensus AUGGCCUGGCGACCGACGGUUCCUUCGAUAAGUCGUUAAGCUAUCGAUGUCUUACUUUCGAUAGUUGGCGGAAGUGCCUUUUUGCGAAGUUACGUUUCUCC______________ ......(((((((...(((.....)))....)))))))(((((((((..........)))))))))((((....))))..................................... (-11.62 = -11.86 + 0.24)



| Location | 8,282,963 – 8,283,053 |
|---|---|
| Length | 90 |
| Sequences | 5 |
| Columns | 115 |
| Reading direction | reverse |
| Mean pairwise identity | 67.38 |
| Shannon entropy | 0.56533 |
| G+C content | 0.44653 |
| Mean single sequence MFE | -23.50 |
| Consensus MFE | -10.74 |
| Energy contribution | -11.22 |
| Covariance contribution | 0.48 |
| Combinations/Pair | 1.26 |
| Mean z-score | -1.49 |
| Structure conservation index | 0.46 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.75 |
| SVM RNA-class probability | 0.807912 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 8282963 90 - 24543557 -------------------------GAAGAAGAAAAAGGCACUUCUGACAACUAUCGAGAGUAAGACAUCGAUAGCGUAACGACUUAUCGAAGGAACCGUCGGUCGUCAGACCAU -------------------------............((....((((((..(((((((..........))))))).....((((...((....))...))))...)))))))).. ( -23.90, z-score = -1.80, R) >droSim1.chr3L 7742235 104 - 22553184 -----------GAUGGAGAAACAAAAACUAGCAAAAAGACACUAUCGCCAACUAUCGAAAGUAAGACAUCGAUAACUUAACGACUUAUCGAAGGAACCGUCGGUCGCCAGGCCAU -----------(((((........................(((.(((........))).)))....(.(((((((.........))))))).)...)))))((((....)))).. ( -20.90, z-score = -1.48, R) >droSec1.super_0 563775 104 - 21120651 -----------GGAGGAGAAACUUAACCUGGCUAAAAGACACUAUCGCCAACUAUCGAAAGUAAGACAUCGAUAACUUAAGGACUUAUCGAAGGAACCGUCGGUCGCCAGGCCAU -----------.(((......)))..((((((.....((((((.(((........))).)))..(((.(((((((.........))))))).(....)))).))))))))).... ( -24.60, z-score = -1.07, R) >droYak2.chr3L 20581374 114 + 24197627 GAAGAAGAAUCUCCAAUUUAACGUAACUUCGCAAAAAGUCACUUCCGCCAACUAUCGAUAGUAAAAU-UCGAUAGCUCGUUAAUGUAUCGAAACUAUCGAGGGUGGCCAGCCCAU ......................((.((((......)))).))....((((.((.((((((((....(-((((((.(........)))))))))))))))).))))))........ ( -25.30, z-score = -1.82, R) >droEre2.scaffold_4784 20008308 102 + 25762168 -------------GAGGAAAAAGAAAGUUCGCAAGAAGGCACUUCCGCCAACUAUCGAAAGUACAACAUCGAUAGCUCGUCAACUUAUCGAAACUAUCGACGGCCGUCAGCCCAU -------------((.((.........(((....)))(((......)))..(((((((..........))))))).)).))......((((.....)))).(((.....)))... ( -22.80, z-score = -1.31, R) >consensus ______________AGAGAAACGAAAACUAGCAAAAAGACACUUCCGCCAACUAUCGAAAGUAAGACAUCGAUAGCUUAACGACUUAUCGAAGGAACCGUCGGUCGCCAGCCCAU .....................................((((((..((........))..)))......(((((((.........))))))).......)))((((....)))).. (-10.74 = -11.22 + 0.48)

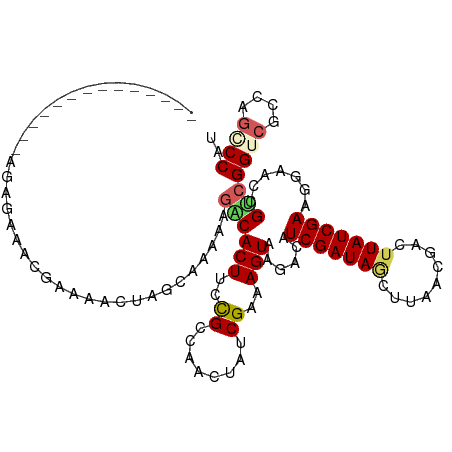

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:11:19 2011