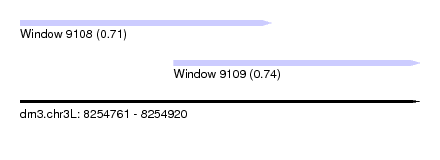
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 8,254,761 – 8,254,920 |
| Length | 159 |
| Max. P | 0.735630 |

| Location | 8,254,761 – 8,254,861 |
|---|---|
| Length | 100 |
| Sequences | 11 |
| Columns | 111 |
| Reading direction | forward |
| Mean pairwise identity | 75.65 |
| Shannon entropy | 0.51574 |
| G+C content | 0.35759 |
| Mean single sequence MFE | -20.78 |
| Consensus MFE | -9.23 |
| Energy contribution | -10.56 |
| Covariance contribution | 1.34 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.69 |
| Structure conservation index | 0.44 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.47 |
| SVM RNA-class probability | 0.710406 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 8254761 100 + 24543557 ---------ACAGUCAAUACGUGAGAUCCUGGC-GAGAUAAAUUCUCUCCCUUAUUUGAAGUUUAUUGUGUGUGAAAAUGCGCUUAAGCUGAAUUUCUAUUAGCC-UUUGU ---------((((......(....).....(((-((((.....))))............((((((..((((((....)))))).))))))............)))-.)))) ( -21.10, z-score = -1.35, R) >droAna3.scaffold_13337 3255828 100 + 23293914 ---------ACAGUCAAUACGUGAGAUCUUUUC-CAGAUAAAUUCUCUCCCUUAUUUGAAGUUUAUUGUGUGUGAAAAUGCGCUUAAGCUGAAUUUCUAUUAGCC-UUUGU ---------((((.........(((((......-(((((((..........))))))).((((((..((((((....)))))).))))))..)))))........-.)))) ( -16.07, z-score = -0.48, R) >droEre2.scaffold_4784 19979125 101 - 25762168 ---------ACAGUCAAUACGUGAGAUGCUAGCCGAGAUAAAUUCUCUCCCUUAUUUGAAGUUUAUUGUGUGUGAAAAUGCGCUUAAGCUGAAUUUCUAUUAGCC-UUUGU ---------((((......(....)..(((((..((((.....))))............((((((..((((((....)))))).)))))).........))))).-.)))) ( -19.80, z-score = -0.93, R) >droYak2.chr3L 20552013 107 - 24197627 --ACAGUGAAUCGUGAGGACGUGAGAUCCUAGC-GAGAUAAAUUCUCUCCCUUAUUUGAAGUUUAUUGUGUGUGAAAAUGCGCUUAAGCUGAAUUUCUAUUAGCC-UUUGU --(((((((((.((.(((((....).)))).))-((((.....)))).............)))))))))((((......))))....(((((.......))))).-..... ( -25.00, z-score = -1.72, R) >droSec1.super_0 535678 100 + 21120651 ---------ACAGUCAAUACGUGAGAUCCUGGC-GAGAUAAAUUCUCUCCCUUAUUUGAAGUUUAUUGUGUGUGAAAAUGCGCUUAAGUUGAACUUCUAUUAGCC-UUUGU ---------((((......(....).....(((-((((.....))))..........((((((((..((((((....))))))......)))))))).....)))-.)))) ( -23.60, z-score = -2.36, R) >droSim1.chr3L 7713440 100 + 22553184 ---------ACAGUCAAUACGUGAGAUCCUGGC-GAGAUAAAUUCUCUCCCUUAUUUGAAGUUUAUUGUGUGUGAAAAUGCGCUUAAGCUGAAUUUCUAUUAGCC-UUUGU ---------((((......(....).....(((-((((.....))))............((((((..((((((....)))))).))))))............)))-.)))) ( -21.10, z-score = -1.35, R) >droWil1.scaffold_180698 2208508 83 + 11422946 ------------GUCAAUAUGUGGAAUUCU---------------UUUUGCUUAUUUGAAGUUUAUUGUGUGUGAAAAUGCGCUUAAGCUGAAUUUCUAUUAGCC-UUUGU ------------.((((...(..(((....---------------)))..)....))))((((((..((((((....)))))).))))))...............-..... ( -16.20, z-score = -1.63, R) >dp4.chrXR_group3a 1407706 89 - 1468910 ------------------ACAUAUGAAAUUCCAUGUGUGUC-UGCUG--GCUUAUUUGAAGUUUAUUGUGUGUGAAAAUGCGCUUAAGCUGAAUUUCUAUUAGCC-UUUGU ------------------(((((((......)))))))...-....(--(((.....((((((((..((((((....))))))......))))))))....))))-..... ( -23.00, z-score = -3.37, R) >droPer1.super_23 1600329 89 - 1662726 ------------------ACAUAUGAAAUUCCAUGUGUGUC-UGCUG--GCUUAUUUGAAGUUUAUUGUGUGUGAAAAUGCGCUUAAGCUGAAUUUCUAUUAGCC-UUUGU ------------------(((((((......)))))))...-....(--(((.....((((((((..((((((....))))))......))))))))....))))-..... ( -23.00, z-score = -3.37, R) >droVir3.scaffold_13049 22683307 110 - 25233164 GAAUCGAAUACGUGAAGAGAUCUUUUUGUUGCGUGUAUCUUGUGCUG-UGCUUAUUUGAAGUUUAUUGUGUGUGAAAAUACGCUUAAGCUGAAUUUCUAUUAGCCCUUUGU .....(((((((..(.((((...)))).)..))))).))....((((-...........((((((..((((((....)))))).))))))..........))))....... ( -22.30, z-score = -1.33, R) >droMoj3.scaffold_6654 1336706 92 + 2564135 ------------------CUGUGUUGUCUGGCAAAUAUCUUGUGCUGCUGCUUAUUUGAAGUUUAUUGUGUGUGAAAAUGCGCUUAAGCUGAAUUUCUAUUAGCC-UUUGU ------------------...........(((.((((......((....))........((((((..((((((....)))))).)))))).......)))).)))-..... ( -17.40, z-score = -0.67, R) >consensus _________ACAGUCAAUACGUGAGAUCCUGGC_GAGAUAAAUUCUCUCCCUUAUUUGAAGUUUAUUGUGUGUGAAAAUGCGCUUAAGCUGAAUUUCUAUUAGCC_UUUGU .........................................................((((((((..((((((....))))))......)))))))).............. ( -9.23 = -10.56 + 1.34)


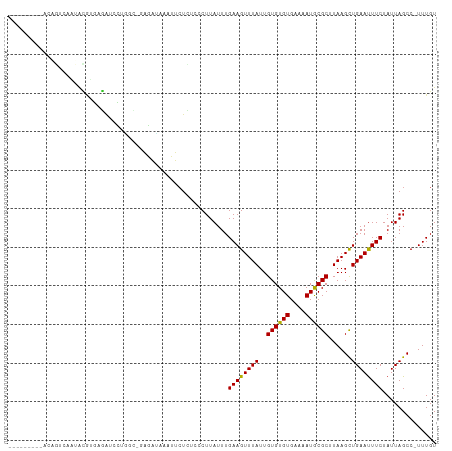
| Location | 8,254,822 – 8,254,920 |
|---|---|
| Length | 98 |
| Sequences | 12 |
| Columns | 117 |
| Reading direction | forward |
| Mean pairwise identity | 79.88 |
| Shannon entropy | 0.41266 |
| G+C content | 0.37333 |
| Mean single sequence MFE | -19.49 |
| Consensus MFE | -9.46 |
| Energy contribution | -9.97 |
| Covariance contribution | 0.51 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.01 |
| Structure conservation index | 0.49 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.54 |
| SVM RNA-class probability | 0.735630 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 8254822 98 + 24543557 GUGAAAAUGCGCUUAAGCUGAAUUUCUAUUAGCCUUUGUG-CGCAUAUUUCACAUCAUCAAUCAAUACGAUAAAAUUUC---ACUCCUUUCAGCCGCUUUUU--------------- ((((((((((((..(((((((.......)))).)))...)-))))).))))))..........................---....................--------------- ( -19.50, z-score = -2.25, R) >droAna3.scaffold_13337 3255889 95 + 23293914 GUGAAAAUGCGCUUAAGCUGAAUUUCUAUUAGCCUUUGUG-CGCAUAUUUCCCAUCAUCAAUCAAUACGAUAAAAGCU-----GCCCCUUUAUCCACCUUU---------------- (.((((((((((..(((((((.......)))).)))...)-))))).)))).)...............((((((....-----.....)))))).......---------------- ( -16.50, z-score = -2.02, R) >droEre2.scaffold_4784 19979187 97 - 25762168 GUGAAAAUGCGCUUAAGCUGAAUUUCUAUUAGCCUUUGUG-CGCAUAUUUCACAUCAUCAAUCAAUACGAUAAAAGUUC---ACUGCUUUCAGCCGUUUUC---------------- ((((((((((((..(((((((.......)))).)))...)-))))).)))))).............(((.(.(((((..---...))))).)..)))....---------------- ( -22.40, z-score = -2.51, R) >droYak2.chr3L 20552081 97 - 24197627 GUGAAAAUGCGCUUAAGCUGAAUUUCUAUUAGCCUUUGUG-CGCAUAUUUCACAUCAUCAAUCAAUACGAUAAAAGUUC---ACUCCUUUCAGCCGUGUUU---------------- ((((((((((((..(((((((.......)))).)))...)-))))).))))))..........((((((.(.((((...---....)))).)..)))))).---------------- ( -23.20, z-score = -3.19, R) >droSec1.super_0 535739 97 + 21120651 GUGAAAAUGCGCUUAAGUUGAACUUCUAUUAGCCUUUGUG-CGCAUAUUUCACAUCAUCAAUCAAUACGAUAAAAGUUC---ACUCCUUUCAGCCGCUUUU---------------- ((((((((((((..(((((((.......)))).)))...)-))))).))))))..................((((((..---.((......))..))))))---------------- ( -17.90, z-score = -1.63, R) >droSim1.chr3L 7713501 97 + 22553184 GUGAAAAUGCGCUUAAGCUGAAUUUCUAUUAGCCUUUGUG-CGCAUAUUUCACAUCAUCAAUCAAUACGAUAAAAGUUC---ACUCCUUUCAGCCGCUUUU---------------- ((((((((((((..(((((((.......)))).)))...)-))))).))))))..................((((((..---.((......))..))))))---------------- ( -20.30, z-score = -2.31, R) >droWil1.scaffold_180698 2208552 94 + 11422946 GUGAAAAUGCGCUUAAGCUGAAUUUCUAUUAGCCUUUGU-UCGCAUAUUUCACAUCAUCAAUCAAUGCGAUAAAAGUU-----UUCCUCCCCGCUC----CCUU------------- (((((((((((..((((((((.......)))))..))).-.))))).)))))).............(((.........-----........)))..----....------------- ( -17.73, z-score = -2.74, R) >dp4.chrXR_group3a 1407756 117 - 1468910 GUGAAAAUGCGCUUAAGCUGAAUUUCUAUUAGCCUUUGUGCCGCAUAUUUCACAUCAUCAAUCAAUACGAUAAAAGUUCGAAAUUCCCCCCAGCUCGGAGUUUUACGUUACCUUUUU ((((((((((((..(((((((.......)))).)))...).))))).)))))).............(((.(((((.(((((.............))))).))))))))......... ( -24.22, z-score = -2.48, R) >droPer1.super_23 1600379 117 - 1662726 GUGAAAAUGCGCUUAAGCUGAAUUUCUAUUAGCCUUUGUGCCGCAUAUUUCACAUCAUCAAUCAACACGAUAAAAGUUCGAAAUUCCCCCCAGUUCGGAGUUUUACGUUACCUUUUU ((((((((((((..(((((((.......)))).)))...).))))).)))))).............(((.(((((.((((((...........)))))).))))))))......... ( -25.20, z-score = -2.91, R) >droVir3.scaffold_13049 22683377 84 - 25233164 GUGAAAAUACGCUUAAGCUGAAUUUCUAUUAGCCCUUUGUGCGCACAUAACACAUCAUCAAUCAAUACGAUAAAAGUUU---AUGCU------------------------------ ..........((.((((((.....((((((.......((((.........)))).........)))).))....)))))---).)).------------------------------ ( -9.09, z-score = 0.83, R) >droMoj3.scaffold_6654 1336759 95 + 2564135 GUGAAAAUGCGCUUAAGCUGAAUUUCUAUUAGCCUUUGUG-CGCAUAUUACACAUCAUCAAUCAAUACGAUAAAAGUUU---AUGCCCGACAAUACGCA------------------ (((((.((((((..(((((((.......)))).)))...)-))))).)).)))......................((.(---((........))).)).------------------ ( -16.90, z-score = -0.54, R) >droGri2.scaffold_15110 21017047 94 - 24565398 GUGAAAAUGCGCUUAAGCUGAAUUUCUAUUAGCCUUUGUGGCGCAUAUUACACAUCAUCAAUCAAUACGAUAAAAGUUG---AUGCC--ACAAAACCCU------------------ (((((.(((((((.(((((((.......)))).)))...))))))).)).)))..(((((((.............))))---)))..--..........------------------ ( -20.92, z-score = -2.39, R) >consensus GUGAAAAUGCGCUUAAGCUGAAUUUCUAUUAGCCUUUGUG_CGCAUAUUUCACAUCAUCAAUCAAUACGAUAAAAGUUC___AUUCCUUUCAGCCCCUUUU________________ (((((.((((((.((((((((.......)))))..))).).))))).)).)))................................................................ ( -9.46 = -9.97 + 0.51)


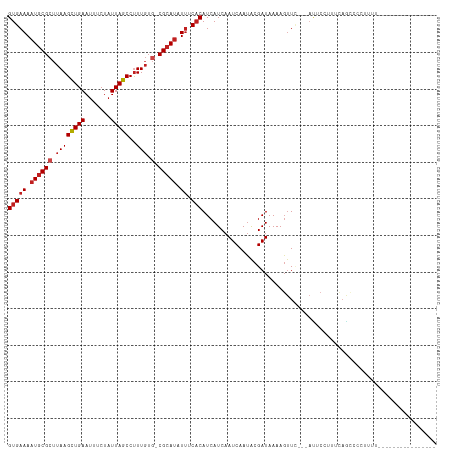
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:11:13 2011