
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 8,158,556 – 8,158,650 |
| Length | 94 |
| Max. P | 0.806048 |

| Location | 8,158,556 – 8,158,650 |
|---|---|
| Length | 94 |
| Sequences | 13 |
| Columns | 114 |
| Reading direction | forward |
| Mean pairwise identity | 75.88 |
| Shannon entropy | 0.49431 |
| G+C content | 0.44078 |
| Mean single sequence MFE | -31.68 |
| Consensus MFE | -11.58 |
| Energy contribution | -11.16 |
| Covariance contribution | -0.42 |
| Combinations/Pair | 1.50 |
| Mean z-score | -2.29 |
| Structure conservation index | 0.37 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.75 |
| SVM RNA-class probability | 0.806048 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 8158556 94 + 24543557 ------UGGGGCUCUGCAC--GUGUUGCCGCGUUUGAAGUUGA---UGCAAAAGUUGUAAAAAAAUUCGCAUCAAUUUUAGUUUUAUACUCAAGAGCAGAGCGCU--------- ------.((.(((((((((--(((....)))))((((((((((---(((.((..((.....))..)).)))))))))))))..............))))))).))--------- ( -33.20, z-score = -3.46, R) >droSim1.chr3L 7624435 94 + 22553184 ------UGGGGCUCUGCAC--GUGUUGCCGCGUUUGAAGUUGA---UGCAAAAGUUGUAAAAAAAUUCGCAUCAAUUUUAGUUUUAUACUCAAGAGCAGAGCGCU--------- ------.((.(((((((((--(((....)))))((((((((((---(((.((..((.....))..)).)))))))))))))..............))))))).))--------- ( -33.20, z-score = -3.46, R) >droSec1.super_0 445526 94 + 21120651 ------UGGGGCUCUGCAC--GUGUUGCCGCGUUUGAAGUUGA---UGCAAAAGUUGUAAAAAAAUUCGCAUCAAUUUUAGUUUUAUACUCAAGAGCAGAGCGCU--------- ------.((.(((((((((--(((....)))))((((((((((---(((.((..((.....))..)).)))))))))))))..............))))))).))--------- ( -33.20, z-score = -3.46, R) >droYak2.chr3L 8747073 94 + 24197627 ------UGGGGCUCUGCAC--GUGUUGCCGCGUUUGAAGUUGA---UGCAAAAGUUGUAAAAAAAUUCGCAUCAAUUUUAGUUUUAUACUCAAGAGCAGAGCGCU--------- ------.((.(((((((((--(((....)))))((((((((((---(((.((..((.....))..)).)))))))))))))..............))))))).))--------- ( -33.20, z-score = -3.46, R) >droEre2.scaffold_4784 22557807 93 - 25762168 ------UGGGGCUCUGCAC--GUGUUGCCGCGUUUGAAGUUGA---UGCAAAAGUUGUAAAAAA-UUCGCAUCAAUUUUAGUUUUAUACUCAAGAGCAGAGCGCU--------- ------.((.(((((((((--(((....)))))((((((((((---(((...((((......))-)).)))))))))))))..............))))))).))--------- ( -33.70, z-score = -3.64, R) >droAna3.scaffold_13337 9841046 96 - 23293914 ------UGGAGCUCUGCAC--GUGUUGCCGCGUUUGAAGUUGA---UACAAAAGUUGUAAAAAAAUUCGCAUCAAUUUUAGUUUUAUACUCG-GUUCGGCGCUCGGCC------ ------..((((...((.(--(....((((.((((((((((((---(.(.((..((.....))..)).).)))))))))))......)).))-)).))))))))....------ ( -20.30, z-score = 0.78, R) >dp4.chrXR_group3a 1199443 101 + 1468910 UGGAGCGUCUGCGGUGCAC--GUGUGGCCGCGUUUGAAGUUGA---UGCAAAAGUUAGACAAAAAUUCGCAUCAAUUUUAGUUUUAUACUCGAGAGCGCGUUGCUC-------- ..(((((((((((.....)--))).)))(((((((((((((((---(((...................)))))))))))(((.....)))...)))))))..))))-------- ( -29.41, z-score = -0.42, R) >droPer1.super_23 1220151 101 + 1662726 UGGAGCGUCUGGGGUGCAC--GUGUGGCCGCGUUUGAAGUUGA---UGCAAAAGUUAGACAAAAAUUCGCAUCAAUUUUAGUUUUAUACUCGAGAGCGCGUUGCUC-------- ....((((((.(((((.((--(((....)))))((((((((((---(((...................))))))))))))).....))))).))).))).......-------- ( -30.31, z-score = -1.07, R) >droWil1.scaffold_180727 138418 95 + 2741493 ------CUGGUCUCUGCACACGUGUUGCCGCGUUUGAAGUUGA---UGCAAAAGUUAGAAAAAA-UUCGCAUCAAUUUUAGUUUUAUACUCAAGAGCAGCCAGCC--------- ------(((((((((....(((((....)))))((((((((((---(((...((((......))-)).)))))))))))))...........))))..)))))..--------- ( -26.10, z-score = -2.06, R) >droVir3.scaffold_13049 7806782 102 + 25233164 -----UGCUAG--CUGCAC--GUGUUGCCGCGUUUGGAGUUGA---UGCAAAAGUUAGAAAAAAUUUCGCAUCAAUUUUAGUUUUAUACUCAAGAGCAGGUCGAGCGCGGCGCC -----(((...--..))).--.....(((((((((((((((((---(((..(((((......))))).))))))))))..(((((.......)))))....))))))))))... ( -33.70, z-score = -1.91, R) >droMoj3.scaffold_6680 11456703 102 + 24764193 -----UGCGAG--CUGCAC--GUGUUGCCGCGUUUGGAGUUGA---UGCAAAAGUUAGAAAAAAUUUCGCAUCAAUUUUAGUUUUAUACUCAAGAGCAGGUCGAGCGCGGCGGC -----(((...--..))).--..((((((((((((((((((((---(((..(((((......))))).))))))))))..(((((.......)))))....))))))))))))) ( -35.50, z-score = -2.22, R) >droGri2.scaffold_15110 6860611 104 + 24565398 -----UGCGAGUGCUGCAC--GUGUUGCCGCGUUUGGAGUUGA---UGCAAAAGUUAGAAAAAAUUUCGCAUCAAUUUUUGUUUUAUACUCAAGAGCAGGUCGAGCGCGGCAAC -----((((.....)))).--..((((((((((((((((((((---(((..(((((......))))).))))))))))(((((((.......)))))))..))))))))))))) ( -38.40, z-score = -3.16, R) >apiMel3.Group6 2044971 94 + 14581788 ------CGUUGUUUUCUCUUGGCGAUUCAGUGUCUUGGGUAGAAAUUGCACAAAUGAGAGAAGA---CGGGUCGAUUC--AUUGCGCAUGGACGCGCAAGGAUCU--------- ------..(((((((((((((......((((.(((.....))).))))......))))))))))---)))...(((((--.((((((......))))))))))).--------- ( -31.60, z-score = -2.17, R) >consensus ______UGGGGCUCUGCAC__GUGUUGCCGCGUUUGAAGUUGA___UGCAAAAGUUAGAAAAAAAUUCGCAUCAAUUUUAGUUUUAUACUCAAGAGCAGGGCGCU_________ .............((((....(((....)))..((((((((((...(((...................)))))))))))))..............))))............... (-11.58 = -11.16 + -0.42)


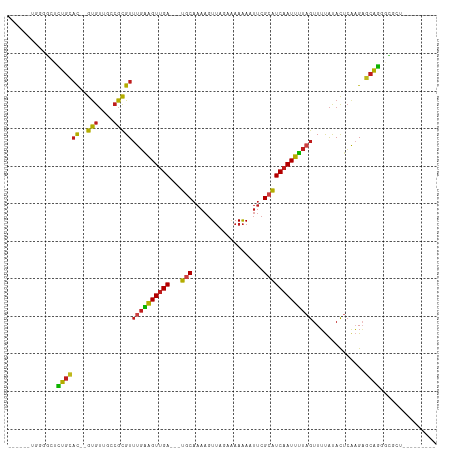
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:11:00 2011