
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 8,055,261 – 8,055,356 |
| Length | 95 |
| Max. P | 0.647535 |

| Location | 8,055,261 – 8,055,356 |
|---|---|
| Length | 95 |
| Sequences | 8 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 62.05 |
| Shannon entropy | 0.70992 |
| G+C content | 0.42127 |
| Mean single sequence MFE | -22.45 |
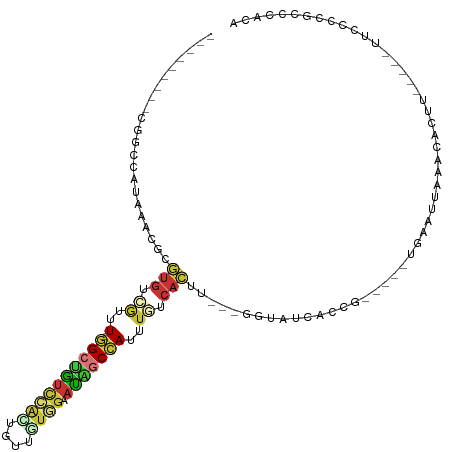
| Consensus MFE | -8.41 |
| Energy contribution | -9.10 |
| Covariance contribution | 0.69 |
| Combinations/Pair | 1.87 |
| Mean z-score | -1.16 |
| Structure conservation index | 0.37 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.33 |
| SVM RNA-class probability | 0.647535 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 8055261 95 - 24543557 ---------CGGCCAUAAACGCGUGUCGUUU-GGCUGUCCACUGUAGUGGAAUAGCCAUUUGUCACUU---GGUAUCACCG-----UGAAUUAAACACUU-----UUCCCAGCCCCCA ---------.(((.........(((.((..(-((((((((((....)))).)))))))..)).)))..---((.....))(-----((.......)))..-----......))).... ( -29.00, z-score = -2.56, R) >droSim1.chr3L 7519197 95 - 22553184 ---------CGGCCAUAAACGCGUGUCGUUU-GGCUGUCCAGUGUUGUGGAAUAGCCAUUUGUCACUU---GGUAUCAGCG-----UGAAUUAAACACUU-----UUCCCCGCCCACA ---------.(((.....(((((((.((..(-(((((((((......))).)))))))..)).))).(---(....)))))-----)(((..........-----)))...))).... ( -26.40, z-score = -1.01, R) >droSec1.super_0 342178 95 - 21120651 ---------CGGCCAUAAACGCGUGUCGUUU-GGCUGUCCAGUGUUGUGGAAUAGCCAUUUGUCACUU---GGUAUCAGCG-----UGAAUUAAACACUU-----UUCCCCGCCCACA ---------.(((.....(((((((.((..(-(((((((((......))).)))))))..)).))).(---(....)))))-----)(((..........-----)))...))).... ( -26.40, z-score = -1.01, R) >droYak2.chr3L 8641937 95 - 24197627 ---------CAGCCAUAAACGCGUGUCGUUU-GGCUGUCCACUGUUGUGGAAUUGCCAUUUAUCACUU---GGUAUCACCG-----UGAAUUAAACACUU-----UUCCCCGCCCACA ---------..........((((((..(..(-(((..(((((....)))))...))))..)..)))..---((.....)))-----))............-----............. ( -22.30, z-score = -0.99, R) >droEre2.scaffold_4784 22452654 95 + 25762168 ---------CAGCCAUAAACGCGUGUCGUUU-GGCUGUCCACUGUUGUGGAAUAGCCAUUUGUCACUU---GGUAUCACCG-----UGAAUUAAACACUU-----UUCCCCGCCCACA ---------..........((((((.((..(-((((((((((....)))).)))))))..)).)))..---((.....)))-----))............-----............. ( -27.40, z-score = -2.35, R) >droAna3.scaffold_13337 9700239 103 + 23293914 ---------CUAACAAAAAUGUAUGUCGUUUAGGCUGUCUG-UGUUUUAGAACGGCCAUUUAUCAUUA---GGAACCAGAGGAAAAUAAUUUAAGCAACU--GCAUCCCCAACCCACC ---------.........(((((.((.((((((((((((((-.....))).)))))).....((.((.---(....).)).))........))))).)))--))))............ ( -18.10, z-score = 0.07, R) >droPer1.super_45 97108 116 + 618639 AAGAGUUUUCCUUCAU-AAUUUAUGAUAUUUUAGCUGUUAAUUGUUAUGGUAUAUCAAUAUAUCAUUUAUGAUAAUCUAAGCUAACAGAUUUAAACCUCUUUGAGUUUUUGACUCAG- (((((.......((((-(...)))))....((((((....((((((((((((((....))))))))....))))))...))))))...........)))))((((((...)))))).- ( -21.40, z-score = -1.64, R) >droGri2.scaffold_15110 12914603 75 + 24565398 -----------------------------UUAUGCCA--AAUUUGGGCAAAUUGUCAACACUUUUUUUAU-GGUACAGAA------UACUUUUAUGAAUU-----UUUUCAGCUCAUU -----------------------------........--....(((((...((((....(((........-)))))))..------........((((..-----..))))))))).. ( -8.60, z-score = 0.22, R) >consensus _________CGGCCAUAAACGCGUGUCGUUU_GGCUGUCCACUGUUGUGGAAUAGCCAUUUGUCACUU___GGUAUCACCG_____UGAAUUAAACACUU_____UUCCCCGCCCACA ......................(((.((....((((((((((....)))).))))))...)).))).................................................... ( -8.41 = -9.10 + 0.69)


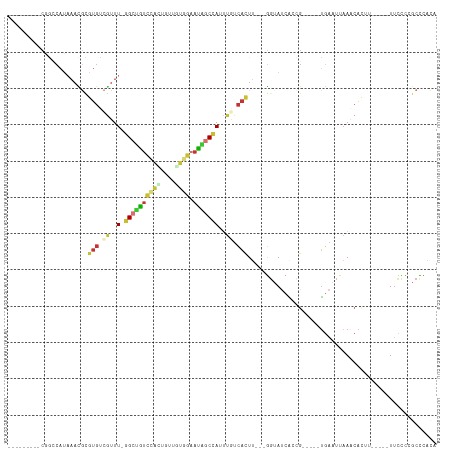
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:10:49 2011