| Sequence ID | dm3.chr3L |
|---|---|
| Location | 7,646,227 – 7,646,347 |
| Length | 120 |
| Max. P | 0.602589 |

| Location | 7,646,227 – 7,646,347 |
|---|---|
| Length | 120 |
| Sequences | 8 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 76.02 |
| Shannon entropy | 0.47395 |
| G+C content | 0.61562 |
| Mean single sequence MFE | -45.94 |
| Consensus MFE | -29.01 |
| Energy contribution | -30.51 |
| Covariance contribution | 1.50 |
| Combinations/Pair | 1.28 |
| Mean z-score | -1.00 |
| Structure conservation index | 0.63 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.23 |
| SVM RNA-class probability | 0.602589 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 7646227 120 - 24543557 AGGAGAUGGAGCGUAAGCACAACCUGCCCCACUGUGUGGGCAAUCUGUUCAUGCGCAGCAUCCAACUCCAAGGCUCUGGGACCGCUUCGAGUUCCGGAGCGGCGAGCAGCGCAUCCGCCG .((((.(((((((((.(.(((...((((((.....).)))))...))).).)))))....)))).))))...((((((((((((...)).))))))))))((((.((...))...)))). ( -51.80, z-score = -1.90, R) >droSim1.chr3L 7160368 120 - 22553184 AGGAGAUGGAGCGUAAGCACAACCUGCCCCACUGUGUGGGCAAUCUGUUCAUGCGCAGCAUCCAACUCCAAGGCUCUGGGACCGGUUCGAGUUCCGGGGCGGCGAGCAGCGCAUCCGCCG .((((.(((((((((.(.(((...((((((.....).)))))...))).).)))))....)))).))))...((((((((((((...)).))))))))))((((.((...))...)))). ( -50.90, z-score = -1.13, R) >droSec1.super_2 7575094 120 - 7591821 AGGAGAUGGAGCGUAAGCACAACCUGCCCCACUGUGUGGGCAAUCUGUUCAUGCGCAGCAUCCAACUCCAAGGCUCUGGGACCGGUUCGAGUUCCGGGGCGGCGAGCAGCGCAUCCGCCG .((((.(((((((((.(.(((...((((((.....).)))))...))).).)))))....)))).))))...((((((((((((...)).))))))))))((((.((...))...)))). ( -50.90, z-score = -1.13, R) >droYak2.chr3L 8252853 120 - 24197627 AGGAGAUGGAGCGCAAGCACAACCUGCCCCACUGUGUGGGCAAUCUGUUCAUGCGUAGCAUCCAACUCCAAGGCUCCGGAACAGGUUCAAGUUCCGGGGCGGCGAGCAGCGCAUCCGCCG .((((.(((((((((.(.(((...((((((.....).)))))...))).).)))))....)))).))))...((((((((((........))))))))))((((.((...))...)))). ( -53.90, z-score = -2.86, R) >droEre2.scaffold_4784 22071039 120 + 25762168 AGGAGAUGGAGCGCAAGCACAACCUGCCCCACUGUGUGGGCAAUCUGUUCAUGCGCAGCAUCCAACUCCAAGGCUCUGGGACCGGUUCGAGUUCCGGGGCGGCGAGCAGCGCAUCCGCCG .((((.(((((((((.(.(((...((((((.....).)))))...))).).)))))....)))).))))...((((((((((((...)).))))))))))((((.((...))...)))). ( -52.90, z-score = -1.43, R) >droAna3.scaffold_13337 16455415 120 - 23293914 AGGAGAUGGAGCGCAAGCACAACCUUCCCCACUGCGUGGGCAACCUGUUCAUGCGCAGCAUCCAGCUCCAGAGCUCGGGAUCGGGUUCCGGUUCGGGAUCAGGAUCCAGUGCAGCAGCAA .((((.((((((....)).............((((((((....)).......))))))..)))).))))...((.(.(((((.(((((((...)))))))..))))).).))........ ( -47.51, z-score = -1.08, R) >dp4.chrXR_group6 1659941 96 - 13314419 CGGACAUGGAGCGCAAGCACAGCCUACCCCAGUGCGUGGGUAACCUCUUCAUGCGCAGCAUUCAGCUGCAGAGCAAUGUGGCCAGCAGUACCACUA------------------------ .((((..((.((....))...((((((........))))))..))......((((((((.....))))).(.((......))).))))).))....------------------------ ( -29.80, z-score = 0.76, R) >droPer1.super_12 2350137 96 - 2414086 CGGACAUGGAGCGCAAGCACAGCCUACCCCAGUGCGUGGGUAACCUCUUCAUGCGCAGCAUUCAGCUGCAGAGCAAUGUGGCCAGCAGUACCACUA------------------------ .((((..((.((....))...((((((........))))))..))......((((((((.....))))).(.((......))).))))).))....------------------------ ( -29.80, z-score = 0.76, R) >consensus AGGAGAUGGAGCGCAAGCACAACCUGCCCCACUGUGUGGGCAAUCUGUUCAUGCGCAGCAUCCAACUCCAAGGCUCUGGGACCGGUUCGAGUUCCGGGGCGGCGAGCAGCGCAUCCGCCG .((((.(((((((((.(.(((...((((((.....).)))))...))).).)))))....)))).)))).....((((((((........)))))))).(((((...........))))) (-29.01 = -30.51 + 1.50)


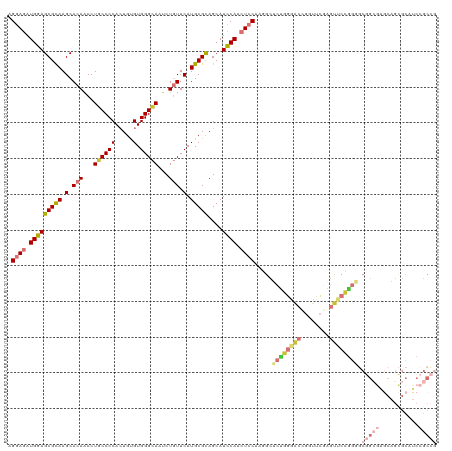
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:10:00 2011